JAX-LM Assignment
The assignment companion for JAX-LM

This assignment serves as a companion for the JAX-LM blog. For the ambitious readers who want to take a stab at implementing our language model in JAX end-to-end, this assignment provides step-by-step instructions and a test suite to guide your implementation.
This assignment is built on top of Assignment 1 (basics): Building a Transformer LM of Stanford’s CS336: Language Modeling From Scratch course.
The main differences are:
- We convert the test cases from PyTorch to JAX
- We add tests and instructions for distributed training in JAX
- Some sections are organized a little differently from the original assignment
However, the modules can mostly be completed in the same order.
Device Requirements
It is strongly recommended to complete this assignment on a system with accelerators (GPU/TPU) to fully observe the speedups from JAX at scale.
That being said, this assignment is compatible with Linux, Windows, and MacOS systems, for sections up to and including Section 3: Training Loop can be completed and run locally on a standard laptop. This will still be helpful for just getting comfortable coding in JAX. However, the distributed implementations assume access to GPUs/TPUs.
If you don’t have acess to accelerators, you can simulate a device mesh using CPUs instead by setting os.environ["XLA_FLAGS"] = "--xla_force_host_platform_device_count=8", where you can replace 8 with your number of desired devices. This must be done before JAX is imported. However, we have not fully tested this configuration.
Getting Started
The blank starter code is provided on the assignment branch. To clone only the starter code:
git clone --single-branch --branch assignment https://github.com/chuyishang/jax-lm.git
Repository Structure The repository is a fork of stanford-cs336/assignment1-basics, and contains:
-
jax_tests/: the core test suite for JAX-based implementations.- Suggested workflow: go through the sections below, passing all non-distributed tests (everything other than
test_sharding). Then go back through to implement sharding. -
adapters.py: This file starts as function stubs. You will edit it as you implement more components to get the relevant tests to run.
- Suggested workflow: go through the sections below, passing all non-distributed tests (everything other than
-
jax_impl/: the directory where your code will live. Feel free to create files and organize your code as you see fit. -
data/tokenizer.pyanddata/train_bpe.py: BPE implementation and training script are provided by default because that code does not differ from the original PyTorch assignment. -
uv.lockandpyproject.tomlwhich provide dependencies including JAX
By default, uv run and uv sync will assume a Linux + CUDA setup and be able to run out of the box. If you are using a CPU-only setup, or on a macOS device, you should first run:
uv sync --no-group cuda
Running Tests Tests can be run using uv run pytest jax_tests/test_file_name.py or uv run pytest -k test_file_name. As a base, you should be able to run
uv run pytest jax_tests/test_train_bpe.py jax_tests/test_tokenizer.py
and pass these tests right away since we’ve provided byte-pair tokenization implementations in data/train_bpe.py and data/tokenizer.py.
1. Basic Building Blocks: NNX Modules
If you’re following along on the CS336 spec, this section is the JAX-equivalent to Section 3: Transformer Language Model Architecture.
In this section, we will build our first nnx.Module classes that will become the building blocks for our Transformer LM. Specifically, we will implement
LinearEmbeddingRMSNormSwiGLU-
RoPE: RoPE does not use random weight initialization
In your implementation, these modules can be defined anywhere under jax_impl/ as long as jax_tests/adapters.py is updated to call them.
Random Number Generation
NNX modules that require random weight initialization should be initialized with an additional rngs parameter. This is because in NNX, random number generation uses explicit states passed in as nnx.Rngs objects.
Thus, your class signature should look something like:
from flax import nnx
import jax.numpy as jnp
class Linear(nnx.Module):
def __init__(self, rngs: nnx.Rngs, in_features: int, out_features: int, dtype: jnp.dtype=jnp.float32):
pass
def __call__(self, x: jnp.ndarray) -> jnp.ndarray:
pass
Another note is that the forward pass of an nnx.Module is implemented directly within the __call__ method, instead of a separate forward() method like in PyTorch.
Device Management
Recall that we also don’t need to pass in devices explicitly. Later on, we will modify this section to support sharding annotations, but we don’t need to worry about that now.
1.1 Linear
Let’s get started with the first module: a linear layer. Again, the class signature for your module should look something like:
class Linear(nnx.Module):
def __init__(self, rngs: nnx.Rngs, in_features: int, out_features: int, dtype: jnp.dtype=jnp.float32):
pass
def __call__(self, x: jnp.ndarray) -> jnp.ndarray:
pass
The Linear module performs the following transformation: \(y = xW\) We first need need to initialize our weight matrix $W \in \mathbb R^{d_{\text{in}} \times d_{\text{out}}}$. Following the original assignment, we will initialize our weights to a truncated normal distribution $W \sim \mathcal N(\mu=0, \sigma^2 = \frac{2}{d_\text{in} + d_\text{out}})$ truncated at $[-3\sigma, 3\sigma]$. This is made easy thanks to nnx.initializers.truncated_normal. You can also play around with more sophisticated initializations offered by the library, although it may not pass the autograder tests.
Once the weight data is initialized, it should be wrapped in an nnx.Param object and stored as a class attribute.
When the function is called, we just apply the linear transformation.
TODOs
- Implement the
Linearmodule using only JAX arrays and methods.
- Note: don’t forget to call the superclass constructor.
- Update
run_linearinadapters.pyto call your method.
- Note: to pass the weights to your linear layer, call
nnx.updatewith the parameters formatted in aStatedictionary. We do not include a bias term in the weights, and your implementation shouldn’t either to work with the tests and adapter as is.- Run
uv run pytest -k test_linearto check against the test case.
If you’re stuck, we provide an in-depth walkthrough for implementing a Linear layer in Implementing Our Model. We only do this for select sections. To just see a completed implementation of all sections, refer to the reference implementation!
1.2 Embedding
Next up is the Embedding module, which takes as input a vector of integer token IDs and converts each of them to their vector representations. Its interface is very similar to Linear:
class Embedding(nnx.Module):
def __init__(self, rngs: nnx.Rngs, n_embeddings: int, embedding_dim: int, dtype: jnp.dtype=jnp.float32):
pass
def __call__(self, token_ids: jnp.ndarray) -> jnp.ndarray:
pass
To make things a bit clearer:
-
n_embeddings= vocabulary size - We maintain a weight matrix $W \in \mathbb R^{\text{n_embeddings} \times \text{embedding_dim}}$ where the $i$-th row gives the embedding for token ID $i$.
- This can be initialized to $W \sim \mathcal N(0, 1)$ truncated at $[-3, 3]$
- The output will itself be a matrix of size $(\text{token_ids.size}, \text{embedding_dim})$
TODOs
- Implement the
Embeddingmodule using only JAX arrays and methods.- Update
run_embeddinginadapters.pyto call your method.- Run
uv run pytest -k test_embeddingto check against the test case.
1.3 RMSNorm
RMSNorm is a basic layer normalization module. For convenience, the formula given a $d$-dimensional vector of activations as input:
class RMSNorm(nnx.Module):
def __init__(self, rngs: nnx.Rngs, d_model: int, eps: float = 1e-5, dtype: jnp.dtype=jnp.float32):
pass
def __call__(self, x: jnp.ndarray) -> jnp.ndarray:
pass
and the formula
\[\text{RMSNorm}(x_{i}) = \frac{x_{i}}{\text{RMS}(x)} g_{i}\]where
\[\text{RMS}(x) = \sqrt{\frac{1}{\text{d\_model}} \sum_{i=1}^{\text{d\_model}} a_{i}^2 + \epsilon}\]Here, our gain vector $g \in \mathbb R^{1 \times \text{d_model}}$ consists of our learnable parameters and $\epsilon$ is a hyperparameter.
TODOs
- Implement the
RMSNormmodule using only JAX arrays and methods.- Update
run_rmsnorminadapters.pyto call your method.- Run
uv run pytest -k test_rmsnormto check against the test case.
1.4 Feed-Forward Network
Now we get into implementing the actual feed-forward network that will be used as part our Transformer LM. The assignment calls for a SwiGLU (Swish-Gated Linear Unit) architecture, which consists of a Gated Linear Unit with a Swish (or SiLU) activation. Concretely:
\[\text{FFN}(x) = \text{SwiGLU}(x, W_1, W_2, W_3) = W_2(\text{SiLU}(W_1 x) \odot W_3 x)\]The dimensions here are:
- $x \in \mathbb R^{d_\text{model}}$
- $W_1, W_3 \in \mathbb{R}^{d_{\text{ff}} \times d_{\text{model}}}$
- $W_2 \in \mathbb{R}^{d_{\text{model}} \times d_{\text{ff}}}$ $d_{\text{ff}}$ is the hidden size of the feed-forward block, while $d_\text{model}$ is the hidden size of the token representation. $d_{\text{ff}}$ should be set to approximately $\frac{8}{3} \cdot d_{\text{model}}$.
The SiLU activation function is \(\text{SiLU}(x) = x \cdot \sigma(x) = \frac{x}{1 + e^{-x}}\)
TODOs
- Implement the SiLU activation function.
- Update
adapters.run_siluand calluv run pytest -k test_siluto test your implementation.- Implement the
SwiGLUmodule using only JAX arrays and methods.
- Note: use your own
Linearclass!- Update
run_swigluinadapters.pyto call your method.- Run
uv run pytest -k test_swigluto check against the test case.
1.5 Relative Positional Embeddings
The last module for this section will implement Rotary Positional Embeddings (RoPE) to inject information about each token’s relative position in the sequence. Here is the class interface:
class RotaryPositionalEmbedding(nnx.Module):
def __init__(self, d_k: int, theta: float, max_seq_len: int):
pass
def __call__(self, x: jnp.ndarray, token_positions: jnp.ndarray) -> jnp.ndarray:
pass
Notably, this is the first module we’ve implemented that doesn’t include rngs in the class signature. This is because this layer has no learnable parameters and thus does not need any randomized initialization.
Because of this, it’s also possible to optimize RoPE by caching the sine and cosine values using nnx.Cache at initialization.
TODO: put the definition of RoPE
TODOs
- Implement the
RotaryPositionalEmbeddingmodule using only JAX arrays and methods.- Update
run_ropeinadapters.pyto call your method.- Run
uv run pytest -k test_ropeto check against the test case.
2. Attention & Full Transformer Block
The modules we have so far together form almost the entire Transformer, with the exception of the actual self-attention part. That’s what we’ll implement now.
In this section, we will implement
- Scaled Dot-Product Attention
- Causal Multi-Head Self-Attention
- Transformer Block
- Transformer Language Model
2.1 Scaled Dot-Product Attention
The SDPA operation takes as input the query, key, and value matrices to perform the attention operation, defined as}”” \(\text{Attention}(Q, K, V) = \text{softmax}\bigg(\frac{Q K^\top}{\sqrt{ d_{k} }} \bigg) \cdot V\) where $Q \in \mathbb R ^{n \times d_{k}}$, $K \in \mathbb R^{m \times d_{k}}$, and $V \in \mathbb R^{m \times d_{v}}$. Here, $m$ is the sequence length and $n$ is the batch size.
For convenience, the numerically-stable softmax activation function is: \(\text{softmax}(x_{i}) = \frac{e^{x_{i} - \max(x)}}{\sum_{j} e^{x_{j} - \max(x)}}\)
TODOs
- Implement the
softmaxactivation function.
- Update
adapters.run_softmaxand calluv run pytest -k test_softmax_matches_pytorchto test your implementation.- Implement the
scaled_dot_product_attentionusing only JAX arrays and methods.
- Note: include an optional
maskparameter which accepts an array of shapeFloat[Array, " ... queries keys"]- Update
run_scaled_dot_product_attentioninadapters.pyto call your method.- Run
uv run pytest -k test_4d_scaled_dot_product_attentionto check against the test case.Hint: if you’ve done the original PyTorch assignment, these methods will look nearly identical.
2.2 Causal Multi-Head Self-Attention
We provide an in-depth walkthrough for implementing Causal Multi-Head Self-Attention in Implementing Our Model.
class MultiHeadSelfAttention(nnx.Module):
def __init__(
self,
rngs: nnx.Rngs,
d_model: int,
num_heads: int,
rope_theta: float = 1e4,
max_seq_len: int = 1024,
):
pass
def __call__(self, x: Array, use_rope: bool = False):
pass
TODOs
- Implement the
MultiHeadSelfAttentionmodule using only JAX arrays and methods.- Update
run_multihead_self_attentioninadapters.pyto call your method.- Run
uv run pytest -k test_multihead_self_attentionto check against the test cases.
2.3 Transformer Block
The last piece of the puzzle is the Transformer block. Conveniently, we’ve already implemented all the submodules we’ll need. Refer to Figure 2 of the below image for the architecture, taken from the original assignment:
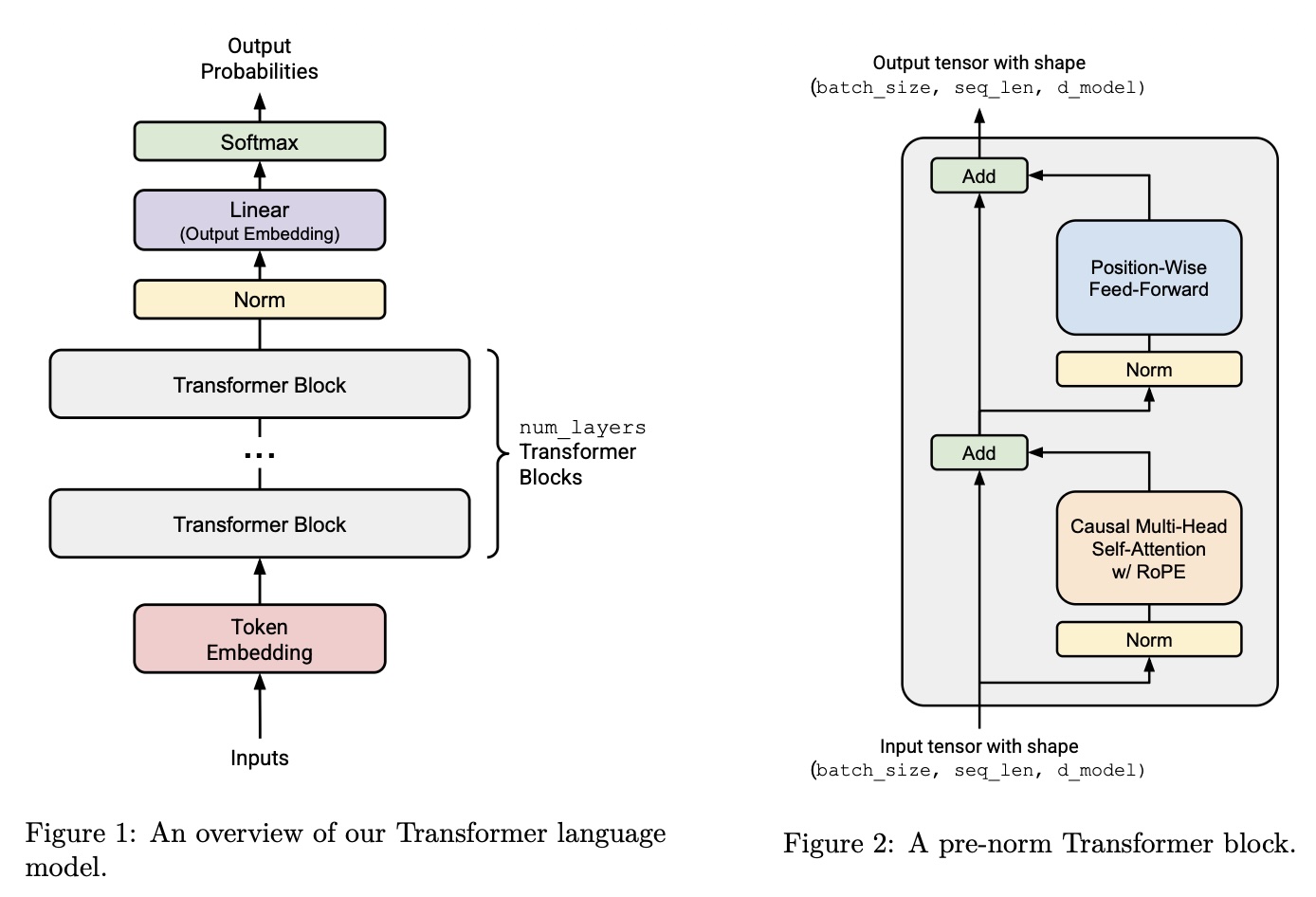
Here’s the TransformerBlock interface:
class TransformerBlock(nnx.Module):
def __init__(
self,
rngs: nnx.Rngs,
d_model: int,
num_heads: int,
d_ff: int,
max_seq_len: int,
theta: float,
):
pass
def __call__(self, x: jnp.ndarray):
pass
TODOs
- Implement the
TransformerBlockmodule using only JAX arrays and methods.- Update
run_transformer_blockinadapters.pyto call your method.- Run
uv run pytest -k test_transformer_blockto check against the test case.
2.4 Transformer LM
Now its finally time to put the pieces together to create the final Transformer language model. The structure is pasted in Figure 1 in the previous subsection. Similar to the Transformer block, we’ve already implemented all the layers we’ll be needing.
class TransformerLM(nnx.Module):
def __init__(
self,
rngs: nnx.Rngs,
d_model: int,
num_heads: int,
d_ff: int,
theta: float,
vocab_size: int,
context_length: int,
num_layers: int,
):
pass
def __call__(self, x: jnp.ndarray):
pass
TODOs
- Implement the
TransformerLMmodule using only JAX arrays and methods.- Update
run_transformer_lminadapters.pyto call your method.- Run
uv run pytest -k test_transformer_lmto check against the test cases.
And there you have it: your own Transformer language model from scratch, completely in JAX. We’ll upgrade this code in Section 4 to add sharding functionality, but this should’ve gotten you comfortable coding in JAX and NNX.
3. Training Loop
Now that we have all the pieces of the model, it’s time to build out the training infrastructure. This will include:
- Cross entropy loss function
- Optax gradient transformations:
- AdamW
- Gradient clipping
- LR schedule
- Train loop with checkpointing
3.1 Cross Entropy Loss
Cross entropy loss is defined as \(\ell_{i}(o, x) = -\log \text{softmax}(o_{i})_{x_{i+1}}\) where the Transformer outputs logits $o \in \mathbb R^{m \times \text{vocab size}}$ for each sequence $x$ of sequence length $m$. Then, the total cross entropy loss for a batch of size $B$ would be \(\mathcal{L}(o, x) = -\frac{1}{B}\sum_{i=1}^{B} \log \big(\mathrm{softmax}(o_i)\big)_{x_i}\) For numerical stability, you should implement the log-sum-exp trick (a helpful blog post about it here).
Note: the test cases can be used as a helpful reference for understanding the expected dimensions of the inputs.
TODOs
- Implement the cross entropy loss function.
- Update
run_cross_entropyinadapters.pyto call your method.- Run
uv run pytest -k test_cross_entropyto check against the test case.Hint: if you’ve done the original PyTorch assignment, these methods will look nearly identical.
3.2 Optimizer
In JAX, the optimizer is treated as a series of Optax transformations. We can first construct our pipeline by adding gradient clipping followed by AdamW. Note that we pass in our learning rate schedule lr_schedule as an argument to AdamW during initialization. Then, we can create an nnx.Optimizer using our model, the pipeline we just created, and the wrt argument that specifies what to optimize.
This pipeline could look something like
def build_optimizer_transform(
optimizer_config: dict,
gradient_clip: float,
lr_schedule: optax.Schedule,
) -> optax.GradientTransformation:
...
TODOs
- Implement the
get_lr_cosine_schedulemethod for cosine annealing LR scheduling using only JAX arrays and methods.- Update
run_get_lr_cosine_scheduleinadapters.pyto call your method.- Run
uv run pytest -k test_get_lr_cosine_scheduleto check against the test case.
3.4 Training Loop
An in-depth walkthrough for is provided in Implementing the Training Loop.
4. Sharding
Now we’ll extend the model and data pipeline to support distributed execution with JAX sharding. The test suite here checks physical sharding (not just logical intent) of:
- input batches,
- model parameters, and
- optimizer states.
We recommend implementing this section after Sections 1-3 are passing.
For the specific sharding specs we want to do for each mode, you can refer to the Implementing Sharding section of the original blog post.
4.1 Running Sharding Tests
The sharding suite is in jax_tests/test_sharding.py. It assumes we have at least 8 devices. If we don’t have that, it simulates 8 devices using CPU with --xla_force_host_platform_device_count=8.
4.2 Mesh + Mode Validation
Sharding is organized around a 2D mesh with axis names data and tensor.
The tests cover four sharding modes: dp, fsdp, tp, and fsdp_tp
The tests also check for valid device meshes for each mode. Invalid mode/mesh combinations should raise ValueError, and unknown mode names should also raise ValueError.
TODOs
- Implement mesh construction (for example via
create_mesh(mesh_shape, mesh_axis_names)), which is called byadapters.get_mesh.- Implement
get_sharding_config_for_mode(mode)and make it raiseValueErrorfor unsupported modes.- Implement
validate_mesh_for_mode(mesh, mode)and enforce valid mode/mesh pairings.- Run:
uv run pytest jax_tests/test_sharding.py -k test_invalid_mode_name_raises -vuv run pytest jax_tests/test_sharding.py -k test_invalid_mode_mesh_combinations_raise -v
4.3 Sharded Batch Pipeline
The sharding tests construct a tiny dataset wrapper with a .data field and call your sharded batch path through adapters. Your implementation should return (inputs, targets) with shape (batch_size, context_length), and both arrays should be sharded according to the mode-specific batch PartitionSpec.
In particular, the tests call:
adapters.get_expected_batch_sharding_spec(mode)adapters.get_sharded_batch(...)
adapters.get_expected_batch_sharding_spec(mode) already forwards to your mode config via:
model.get_batch_sharding_for_mode(mode)
adapters.get_sharded_batch(...) is still a TODO adapter path and should route to your sharded training/data batch utility.
TODOs
- Implement the batch sharding policy for each mode (e.g. via
get_batch_sharding_for_mode(mode)).- Implement sharded batch loading/sampling and ensure returned arrays are placed with the requested
PartitionSpec.- Implement
adapters.get_sharded_batch.- Run
uv run pytest jax_tests/test_sharding.py -k test_batch_sharding -v.
4.4 Sharded Model + Optimizer Initialization
Next, shard model parameters and optimizer state. The tests call your model/optimizer initialization path in both:
- sharded mode (
mesh+ sharding config), and - no-sharding mode (
mesh=None,sharding_config=None).
No-sharding mode should still initialize correctly and keep arrays local (single addressable shard per array). The sharding tests also check that the optimizer state arrays are physically sharded.
TODOs
- Implement
create_model_and_optimizer(rngs, model_config, optimizer_config, sharding_config, mesh)so it supports both sharded and unsharded paths.- Make sure optimizer states are initialized with sharding consistent with their corresponding parameters.
- Implement
adapters.get_sharded_model_and_optimizer.- Run:
uv run pytest jax_tests/test_sharding.py -k test_model_sharding -vuv run pytest jax_tests/test_sharding.py -k test_optimizer_sharding -vuv run pytest jax_tests/test_sharding.py -k test_no_sharding_still_supported -v
Once this section passes, your implementation supports both local and distributed initialization paths and validates that data/model/optimizer arrays are actually distributed across devices.
Congratulations on finishing this assignment!
Enjoy Reading This Article?
Here are some more articles you might like to read next: