JAX-LM: Language Modelling and Distributed Training in JAX
A guide to language modelling and distributed training from scratch in JAX
Introduction
In this blog, we’ll implement a language model from scratch in JAX, then scale it up with distributed training across multiple GPUs/TPUs.
We draw heavily upon 2 great resources. The first is Assignment 1 from Stanford’s CS336 course, which goes over how to implement language models from scratch in PyTorch. The second resource is the How to Scale Your Model textbook, which explains the theory behind parallelism methods and distributed training.
The goal of this blog is to connect these two perspectives by:
- Implementing a language model from scratch in JAX instead of PyTorch
- Extending this implementation with distributed training in both JAX and PyTorch
- Providing a simple, customizable codebase for empirically validating theoretical compute rooflines.
We also provide an assignment with test cases so you can implement all of this in JAX yourself!
The associated code can be found at https://github.com/chuyishang/jax-lm/. If you want to try coding it yourself, you can use the
assignmentbranch of the repo as a starter. The code follows the same flow as CS336 Assignment 1, but with custom test cases for JAX and distributed training.
This blog is split into 4 parts. First, we will provide a brief introduction to JAX and NNX. Next, we will implement the basic components of a language model in JAX, while providing comparisons to PyTorch. Then, we will introduce various distributed training methods (DP, TP, FSDP, FSDP+TP) and show how to implement them in JAX. Finally, we will use our code to empirically validate theoretical compute rooflines (coming soon!).
With that being said, let’s dive in!
Preliminaries
How to Use this Resource
This post is designed to be read front-to-back, but feel free to jump to whatever is most relevant to you. If you’re already comfortable with JAX and NNX, you can go straight to Implementing Our Model. If you’re familiar with distributed training concepts (DP, FSDP, TP) and just want to see the JAX implementation, skip ahead to Implementing Distributed Training and the code implementation.
In addition to this blog post, we also release:
- An assignment for you to implement a JAX-based language model from scratch. This is based on Assignment 1 of Stanford’s CS336 course, but we add tests in JAX and tests for distributed training (DP, FSDP, TP, FSDP+TP). The assignment can be found here, and the starter code can be found here.
- We provide code implementations of a simple, non-distributed language model in both JAX and PyTorch. You can compare these two implementations side-by-side to get a feel of the differences. These can be found here:
- We extend both the JAX and PyTorch implementations to support distributed training with DP, FSDP, TP, and combined FSDP+TP. These can be found here:
This blog post is a living document and may contain errors. If you spot anything, please leave a comment below!
Recommended Background
As mentioned, this blog post borrows a lot of code from the CS336 Assignment 1, an excellent resource for learning how to build a language model from scratch with PyTorch. If you feel shaky on your PyTorch or Language Modeling fundamentals, we recommend you start there.
This blog post also references the How to Scale Your Model book, which describes distributed training and sharding with JAX in much more detail. We recommend reading that book in parallel with this blog post to gain a deeper understanding of distributed training.
With that being said, let’s dive in!
Intro to JAX and NNX
So what is JAX, and why would we want to use it at all? In this section, we’ll try to answer these questions by providing a brief overview of JAX and its neural network API, NNX. We will also touch upon some key concepts we’ll need for implementing our language model from scratch.
Note on Documentation
This section is not meant to be a comprehensive guide or a replacement for the docs. We recommend referring to the JAX and Flax docs for a full reference.
Intro to JAX
JAX is an open-source Python library for high-performance numerical computing developed by Google. It combines a NumPy-like syntax with the ability to run on accelerators (GPUs/TPUs), and also provides an automatic differentiation system for efficiently computing gradients. While JAX was initially developed to be compatible with Google’s Tensor Processing Units (TPUs), it is also compatible with GPUs and CPUs and can provide significant speedups on these devices.
So what makes JAX unique?
-
Firstly, JAX uses a functional programming model. Unlike PyTorch’s stateful, object-oriented approach, JAX adopts pure functions and immutable arrays. As an example, there is no model object that holds parameters. Instead, in core JAX, the model is represented as a nested collection of arrays (a PyTree) and passed explicitly into functions. This makes the execution model more transparent and composable.
- Secondly, JAX provides a set of very powerful composable function transformations. The core idea is that we can write a plain Python or NumPy function and transform it into a more powerful function. Some common transforms are as follows:
-
jax.grad, which differentiates a function -
jax.jit, which compiles a function via XLA for fast execution -
jax.vmap, which vectorizes a function over some batch dimension automatically -
jax.pmap, which is similar tojax.vmapbut parallelizes a function across devices These functions can be composed arbitrarily (such asjit(vmap(grad(f)))) and can often provide huge speedups in practice.
-
- Lastly, JAX uses the XLA Compiler as its backend, which optimizes across the entire computation graph instead of operator by operator as PyTorch does. This allows XLA to perform aggressive fusion and optimizations that span multiple operations and even devices, which can unlock large performance gains.
Putting this all together, the use of pure, composable functions allows the JAX compiler to trace entire functional units into an intermediate representation, called the jaxpr, instead of executing each line in the Python interpreter step-by-step. That’s where the speedup comes from.
Let’s now take a look at some of the code behind these function transformations!
jax.grad
At the heart of any deep learning library is automatic differentiation (autodiff), and JAX’s approach is built around jax.grad. jax.grad is a transform that takes in a function f and returns a new function that computes its gradient.
This is a slightly different mental model from PyTorch. In PyTorch, the computation graph is built incrementally as we perform each operation in the forward pass. To get the backwards pass computation, PyTorch walks through the graph in reverse.
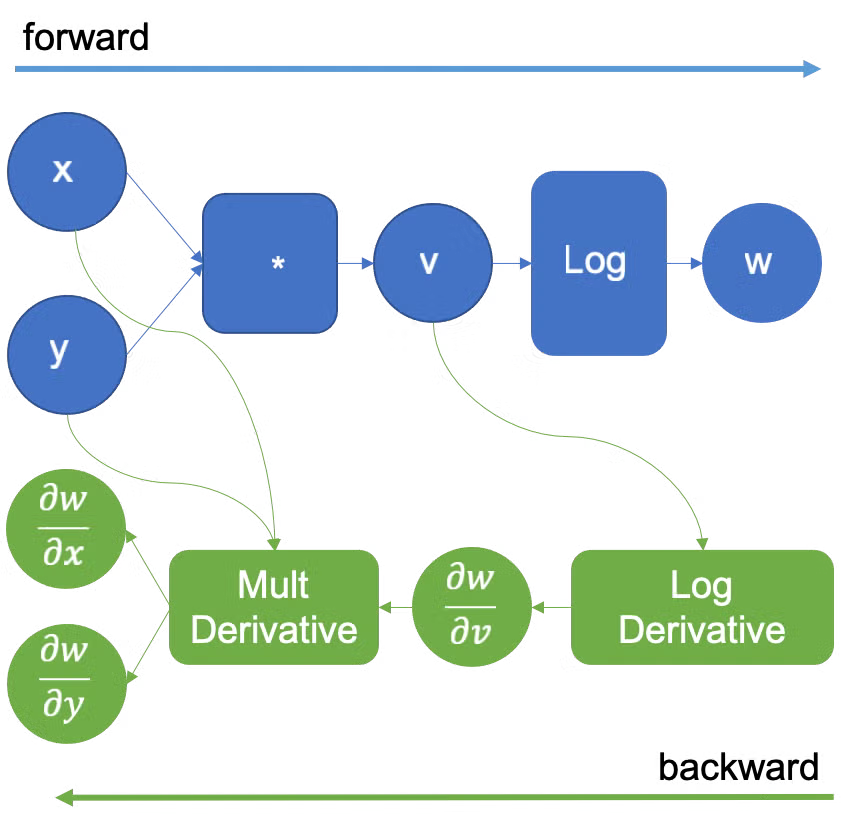
On the other hand, differentiation in JAX is expressed directly as a function transformation. If f computes a scalar output, then jax.grad(f) is a new function with the same inputs that returns the gradient of f with respect to whichever arguments we specify.
Code
To see this in code form, let’s look at the following example of a simple loss function:
import jax
import jax.numpy as jnp
def loss(params, x, y):
pred = params @ x
return jnp.mean((pred - y) ** 2)
grad_fn = jax.grad(loss, argnums=(0,1))
grads = grad_fn(params, x, y)
dparams, dx = grads # for clarity
We can define our loss function normally, using the NumPy-like operation jnp.mean to take the mean of our vector. Then, we can call jax.grad on our loss function to transform it into a new function grad_fn that computes the gradients of its inputs.
One thing to note is that the new function grad_fn will be a wrapper around the original loss function. When we call grad_fn , JAX will run the forward pass (loss) and backwards pass (getting the gradients) in the same call. As a result, grad_fn must have the same function signature as the original loss function.
To specify which arguments to differentiate w.r.t., we can pass in argnums. Here, argnums=(0,1) corresponds to params and x, so grad_fn returns a tuple of 2 gradients, dparams and dx.
If we look carefully, the last two lines can actually be combined into the following form:
grads = jax.grad(loss)(params, x, y)where jax.grad acts as a higher order function. If you are familiar with Python decorators, we see that this is actually the exact pattern of a decorator! So an equivalent way of writing the entire code would just be:
import jax
import jax.numpy as jnp
@jax.grad
def loss(params, x, y):
pred = params @ x
return jnp.mean((pred - y) ** 2)
grads = loss(params, x, y)Since all JAX transforms are higher-order functions of the same form, they can all be used as decorators in this way.
But while this is a nice illustration of the JAX programming model, for gradients specifically this is usually not the most convenient form. This is because after decoration with @jax.grad, our loss function would only return the gradients instead of the actual loss!
Since we need both the loss in addition to the gradients during training, we typically use the jax.value_and_grad transform instead. The usage is identical, except the returned function now returns a (loss, grads) tuple instead of just grads.
loss_and_grad_fn = jax.value_and_grad(loss)
loss_value, grads = loss_and_grad_fn(params, x, y)jax.jit
jax.jit is a transform that compiles a function using the XLA compiler.
JIT compilation is powerful because XLA sees the whole function and optimizes it its entirety. This allows it to fuse operations, optimize memory layout, and parallelize in ways that line-by-line execution cannot.
So what actually happens when we compile a function? On the first call, JAX traces the function by running it with abstract placeholders. You can think of these as something similar to types: instead of passing in x = jnp.array([1.0, 2.0, 3.0]), JAX traces with something like x = float32[3]. The idea is that XLA doesn’t need to know exact values of arrays to build the computation graph for optimization, it only needs to know the shapes and types of data.
After tracing the function to construct a computation graph, JAX compiles it to optimized XLA bytecode. On subsequent calls, we can just run the compiled binaries instead of the Python code.
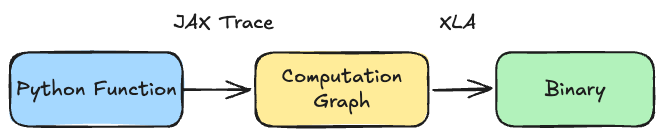
In practice, this can speed up code by a lot. The speedup usually gets larger the more operations we fuse in one jax.jit call, which is why many implementations wrap the entire training step (forward + backward + optimizer update) into a single jax.jit block, which we show below.
Additional Notes on Compilation
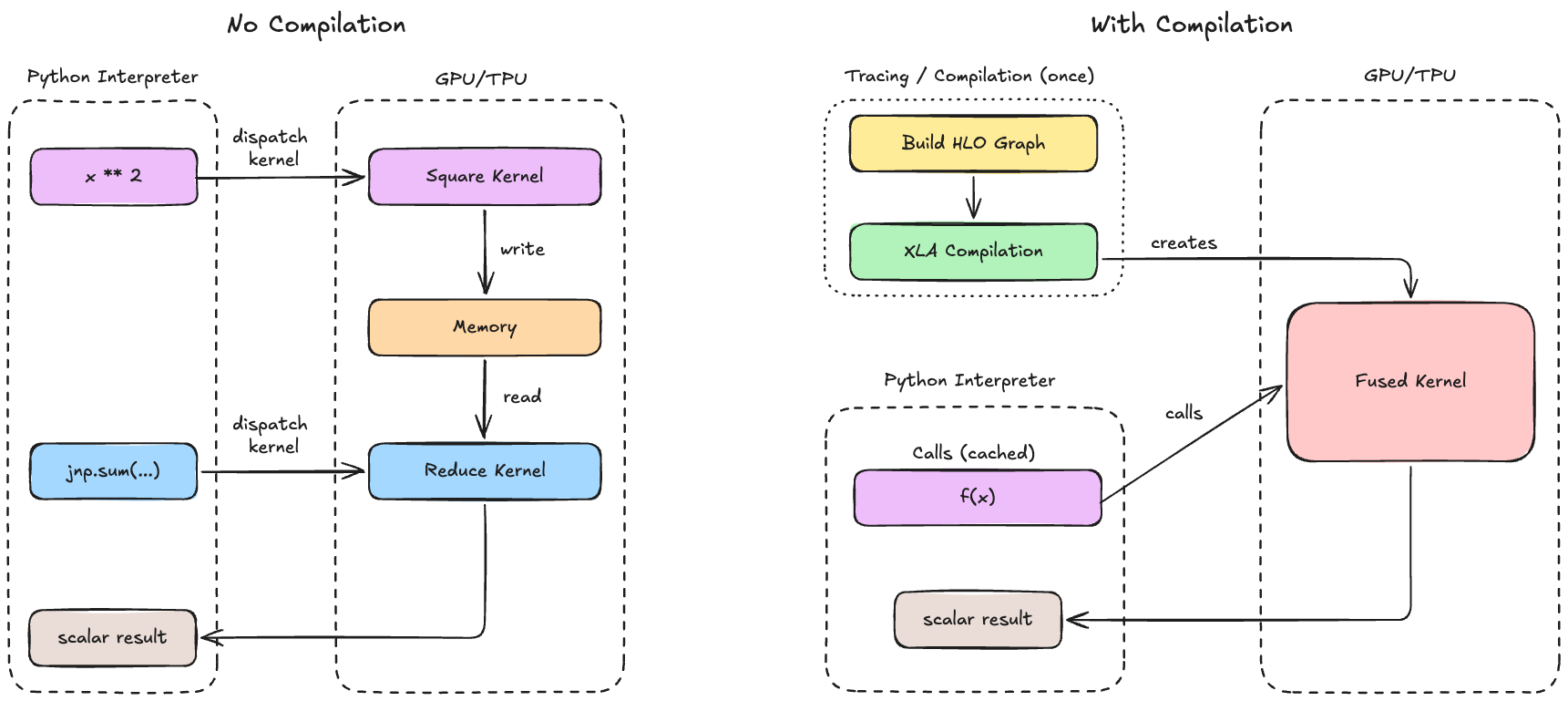
Why do we want to compile? Python can be slow for numerical computation since each operation needs to go through the Python interpreter, which has significant overhead. For example, performing jnp.sum(x ** 2) actually requires 2 operations (** and sum), which are dispatched separately to the GPU/TPU.
Thus, each operation launches its own kernel and must read/write intermediate results from memory, which are both quite expensive. Compilation solves this by seeing the entire computation graph at once, allowing us to merge multiple operations into a single GPU kernel, reorder operations to maximize cache usage, and precompute anything that doesn’t depend on inputs (and store it for future calls).
Code
Here is what jax.jit might look like in code form:
@jax.jit
def f(x):
return jnp.sum(x ** 2)
x = jnp.ones((3,))
output = f(x)A more realistic training step might look something like this:
@jax.jit
def train_step(params, x, y):
loss, grads = jax.value_and_grad(loss_fn)(params, x, y)
params = params - lr * grads
return params, loss
for batch_x, batch_y in data:
params, loss = train_step(params, batch_x, batch_y)Note that we do the entire forwards, backwards, and parameter updates in a single function compiled by jax.jit!
Common Pitfalls
Note that JAX recompiles functions if the shapes of the input change. Take a look at the following example:
@jax.jit
def f(x):
return jnp.sum(x ** 2)
f(jnp.ones((3,))) # compiles for float32[3]
f(jnp.zeros((3,))) # uses cached compiled function since same shape
f(jnp.ones((5,))) # !! recompiles for float32[5] !!
f(jnp.ones((7,))) # !! recompiles for float32[7] !!
f(jnp.zeros((3,))) # uses cached compiled function from the first callJAX compilation takes into account the input shapes to best optimize the program, so it saves a separate binary per shape. In our example, JAX compiles f for the first time with a float32[3] input and can reuse this for the second call. However, we trigger a costly recompilation each time our input shape is different!
Secondly, since JAX traces Python control flow at compile time, ordinary if statements and loops can only depend on values known at trace time. Static loops are fine, but if a branch depends on a runtime value, a Python if inside jax.jit will fail. In those cases, we should use JAX control-flow primitives such as lax.cond, lax.scan, lax.fori_loop, or lax.while_loop.
For example, the following looks natural in Python, but fails under jax.jit because the branch depends on the runtime value of x:
import jax
@jax.jit
def bad_f(x):
count = 0
for _ in range(3):
count += 1 # static loop: OK
if x > 0: # not OK: value-dependent Python branch under jit
return x + count
else:
return -x + count
# bad_f(1.0) # raises TracerBoolConversionErrorTo express the same logic in a JIT-compatible way, we can replace the Python if with lax.cond, which takes a scalar predicate and two branch functions:
import jax
from jax import lax
@jax.jit
def f(x):
count = 0
for _ in range(3):
count += 1 # static loop: OK
return lax.cond(
x > 0,
lambda x: x + count,
lambda x: -x + count,
x,
)Finally, any Python-specific side effects (such as print statements) only happen at trace time and not during executing the compiled program. As a result, a print statement inside a JIT-compiled function may appear only when JAX first traces the function and on retraces. Here’s what this might look like:
@jax.jit
def f(x):
print(x)
>>> f(1)
JitTracer<~int32[]>
>>> f(2) # same shape, no print since we use cached compiled function
>>> f(jnp.array([3, 4])) # different shape, retrace
JitTracer<int32[2]>
Note that these prints happen only at trace time, so JAX does not yet know the concrete value, only the abstract tracer type!
Tip for Print Debugging
To print in a JIT-compiled function, we can use jax.debug.print instead of the default print
jax.vmap
The final core JAX transforms are for vectorization and parallelization. jax.vmap stands for “vectorized map” and transforms a function written for a single example into one that operates over a batch.
import jax
import jax.numpy as jnp
def f(x):
return jnp.dot(x, x)
batched_f = jax.vmap(f)
batched_f(jnp.ones((8, 4)))Equivalently, we can also have this in decorator form:
@jax.vmap
def f(x):
return jnp.dot(x, x)
f(jnp.ones((8, 4)))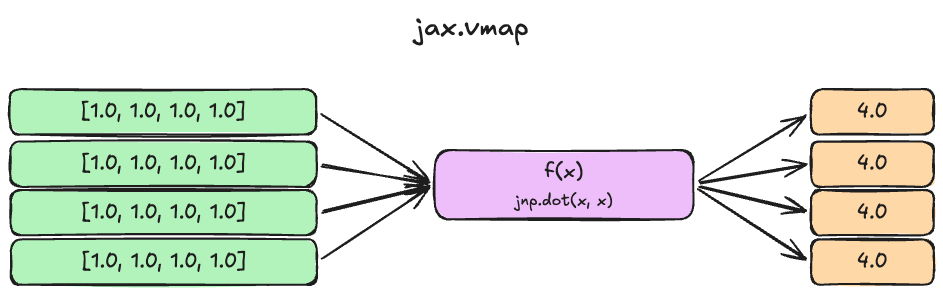
An illustration of the above jax.vmapcode.
The idea here is that we can define a dot product function f that operates on a single vector, but then use jax.vmap to make it batched. By default, jax.vmap maps over the first dimension, but we can also specify which dimensions we want to batch over by passing in the in_axes argument.
jax.pmap
jax.pmap stands for parallel map. It’s similar to vmap but operates over devices (different GPUs/TPUs) by running each slice of the batch on a separate accelerator. We describe pmap for background and historical completeness, but it’s worth noting that newer approaches often prefer mesh-based sharding instead, which we’ll discuss later in the parallelism section.
n_devices = jax.device_count()
def loss_fn(params, x):
return jnp.sum(params * x)
parallel_loss = jax.pmap(loss_fn, in_axes=(None, 0))
xs = jnp.ones((n_devices, 4)) # one slice per device. note that the batch dimension must equal `n_devices`
parallel_loss(params, xs) # runs shard i on device iIn this example, we pass in in_axes=(None, 0). This means that we want the first argument params to replicated across all devices (None), and the second argument to be split across the device axis 0. Since we don’t specify a device mesh, our devices form a single (n_devices,) mesh, so this effectively sends row i to device i.
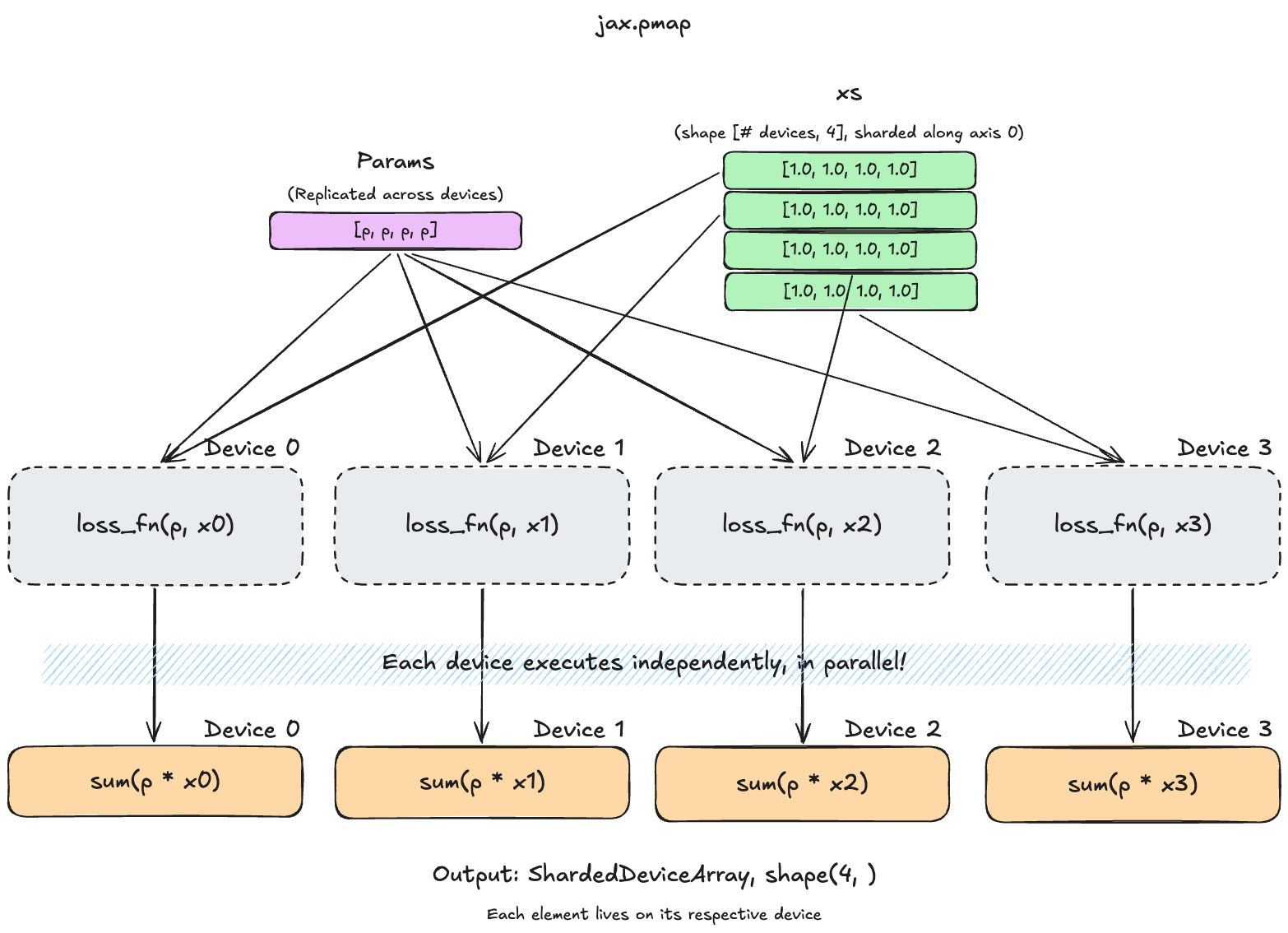
PyTorch comparison for jax.pmap
The equivalent code in PyTorch would look something like the following:
import torch
import torch.distributed as dist
def loss_fn(params, x):
return torch.sum(params * x)
def main():
dist.init_process_group(backend='nccl')
rank = dist.get_rank()
world_size = dist.get_world_size()
params = torch.ones(4).cuda(rank)
xs = torch.ones((world_size, 4)).cuda(rank) # full batch, same shape as JAX
x_local = xs[rank] # manually shard: each process takes its slice
loss = loss_fn(params, x_local)
print(f"rank {rank}, loss: {loss}")
if __name__ == "__main__":
main()and we would need to launch it with torchrun --nproc_per_node=4 script.py. As you can see, we would need to manually send shards to their assigned devices and each process would also run its own Python interpreter. Meanwhile, JAX allows us to run all of this in a single process with pmap and provides cleaner scalability for more complex distributed methods.
NNX
Now that we’ve talked about some of the core concepts behind JAX, let’s talk a bit about NNX, which is what we will be using for the rest of this blog.
NNX is the neural network API for JAX. If you are familiar with PyTorch, you can think of NNX as playing a role similar to torch.nn. NNX was introduced in 2024 as an evolution of Flax Linen, with the goal of keeping the parts of Linen that worked well while making the programming model more Pythonic and easier to debug.
In particular, NNX relaxes the purely functional approach of core JAX by introducing a module-based, object-oriented design similar to PyTorch. Instead of passing model states explicitly in and out of functions, NNX now allows us to define stateful module objects. NNX then provides object-aware versions of common JAX transforms so we can still leverage JAX’s power within this more familiar programming model. For example, @jax.jit becomes @nnx.jit, and jax.value_and_grad becomes nnx.value_and_grad.
To learn a bit more about why you should use NNX, you can refer to the official explanation here.
Implementing Our Model
In this section, we will implement a basic version of a Transformer language model in JAX and provide a version of the same code written in PyTorch for comparison. We will begin with a simple version without worrying too much about sharding and distributed training, then layer in that functionality afterwards.
Let’s start by implementing the simplest component of any modern day neural network: the Linear layer.
The Linear Layer
The following is how you might implement a linear layer (without biases) in PyTorch:
class Linear(nn.Module):
def __init__(
self,
in_features: int,
out_features: int,
device: torch.device | None = None,
dtype: torch.dtype | None = None
):
super().__init__()
weights = torch.empty(out_features, in_features, device=device, dtype=dtype)
std = (2 / (in_features + out_features)) ** 0.5
nn.init.trunc_normal_(weights, mean=0.0, std=std, a=-3.0 * std, b=3.0 * std)
self.weights = nn.Parameter(weights)
def forward(self, x: Tensor) -> Tensor:
return einsum(x, self.weights, "... d_in, d_out d_in -> ... d_out")And here is how you would do it in NNX:
class Linear(nnx.Module):
def __init__( self,
rngs: nnx.Rngs,
in_features: int,
out_features: int,
dtype: jnp.dtype = jnp.float32,
):
super().__init__()
std = (2 / (in_features + out_features)) ** 0.5
init_fn = nnx.initializers.truncated_normal(stddev=std, lower=-3.0 * std, upper=3.0 * std)
weights_data = init_fn(rngs.params(), (in_features, out_features), dtype)
self.weights = nnx.Param(weights_data)
def __call__(self, x: jnp.ndarray) -> jnp.ndarray:
return jnp.einsum("...i,io->...o", x, self.weights.get_value())Weight Storage Convention
In PyTorch, the weights for a Linear layer are typically stored with shape [Out, In], whereas in JAX, the weights are stored as [In, Out].
As you can see, the code looks very similar. Aside from minor differences in syntax and storage convention, there are a few key changes that illustrate some of the philosophical differences between JAX and PyTorch.
-
Initialization: In PyTorch, initialization functions usually operate in-place on an already-allocated tensor. This is why we create an empty weights tensor in the desired shape and on the desired device before initializing it. On the other hand, initialization functions in JAX typically return a tensor (
weights_datain this case), which we then wrap withnnx.Param()to make it a parameter object so that JAX knows how to use it for transforms and state management. The actual weight tensor of annnx.Paramactually lives in its.valueattribute.This example highlights a core philosophical difference between PyTorch and JAX. PyTorch is designed in an object-oriented style centered around stateful modules, where parameters live inside modules and workflows mutate tensors in place. JAX treats parameters as explicit data and often represents common workflows as functions that produce new values.
-
Device Placement: In PyTorch, device placement is explicit. Typically, we create parameters directly on a target device with
device=...or move them afterwards with.to(device). Meanwhile, in JAX, device placement is implicit and determined automatically by the compiler and any sharding configs we specify. -
RNG: In JAX, RNGs need to be passed in explicitly, and are split/consumed deterministically. On the other hand, RNG is largely global and ambient in PyTorch (i.e. we can set a global RNG seed) and we don’t need to pass it explicitly to each function.
In the example above, we use
rngs.params()in theinit_fn, which generates an random number from theparamsstream. There are 2 main PRNG key stream names used by Flax NNX’s built-in layers, which areparamsanddropout.
Some other minor differences:
- PyTorch uses the
forwardconvention, which is wrapped by the__call__method implemented by the parent classnn.Module. JAX typically uses__call__directly.
Multi-head Self Attention
Let’s take a look at another example before moving on. For those following along on the assignment, note that the other building blocks of our model have minor syntax differences compared to PyTorch, but follow the same ideas discussed here.
This is what code for multi-head self attention (MHA) in PyTorch would look like:
class MultiHeadSelfAttention(nn.Module):
def __init__(
self,
d_model: int,
num_heads: int,
rope_theta: float = 1e4,
max_seq_len: int = 1024,
device: torch.device | None = None,
dtype: torch.dtype | None = None,
):
super().__init__()
assert d_model % num_heads == 0, "d_model must be divisible by num_heads"
self.num_heads = num_heads
self.d_k = d_model // num_heads
self.Q_proj = Linear(d_model, d_model, device=device, dtype=dtype)
self.K_proj = Linear(d_model, d_model, device=device, dtype=dtype)
self.V_proj = Linear(d_model, d_model, device=device, dtype=dtype)
self.O_proj = Linear(d_model, d_model, device=device, dtype=dtype)
self.register_buffer(
"causal_mask",
torch.tril(torch.ones(max_seq_len, max_seq_len, device=device)),
persistent=False,
)
self.rope = RotaryPositionalEmbedding(self.d_k, theta=rope_theta, max_seq_len=max_seq_len, device=device)
def forward(self, x: Tensor, use_rope: bool = False) -> Tensor:
batch_size, seq_len, _ = x.shape
q = self.Q_proj(x).reshape(batch_size, seq_len, self.num_heads, self.d_k).transpose(1, 2)
k = self.K_proj(x).reshape(batch_size, seq_len, self.num_heads, self.d_k).transpose(1, 2)
v = self.V_proj(x).reshape(batch_size, seq_len, self.num_heads, self.d_k).transpose(1, 2)
if use_rope:
positions = torch.arange(seq_len, device=x.device)
q = self.rope(q, positions)
k = self.rope(k, positions)
attn_out = sdpa(q, k, v, self.causal_mask[:seq_len, :seq_len])
attn_out = attn_out.transpose(1, 2).contiguous().reshape_as(x)
return self.O_proj(attn_out)And here is what it looks like in JAX:
class MultiHeadSelfAttention(nnx.Module):
def __init__(
self,
rngs: nnx.Rngs,
d_model: int,
num_heads: int,
rope_theta: float = 1e4,
max_seq_len: int = 1024,
dtype: jnp.dtype = jnp.float32,
):
super().__init__()
assert d_model % num_heads == 0, "d_model must be divisible by num_heads"
self.num_heads = num_heads
self.d_k = d_model // num_heads
self.Q_proj = Linear(rngs=rngs, in_features=d_model, out_features=d_model, dtype=dtype)
self.K_proj = Linear(rngs=rngs, in_features=d_model, out_features=d_model, dtype=dtype)
self.V_proj = Linear(rngs=rngs, in_features=d_model, out_features=d_model, dtype=dtype)
self.O_proj = Linear(rngs=rngs, in_features=d_model, out_features=d_model, dtype=dtype)
self.causal_mask = jnp.tril(jnp.ones((max_seq_len, max_seq_len)))
self.rope = RotaryPositionalEmbedding(d_k=self.d_k, theta=rope_theta, max_seq_len=max_seq_len)
def __call__(self, x: Array, use_rope: bool = False):
batch_size, seq_len, _ = x.shape
q = self.Q_proj(x).reshape(batch_size, seq_len, self.num_heads, self.d_k).swapaxes(1, 2)
k = self.K_proj(x).reshape(batch_size, seq_len, self.num_heads, self.d_k).swapaxes(1, 2)
v = self.V_proj(x).reshape(batch_size, seq_len, self.num_heads, self.d_k).swapaxes(1, 2)
if use_rope:
positions = jnp.arange(seq_len)
q = self.rope(q, positions)
k = self.rope(k, positions)
attn_out = sdpa(q, k, v, self.causal_mask[:seq_len, :seq_len])
attn_out = attn_out.swapaxes(1, 2).reshape(x.shape)
return self.O_proj(attn_out)Aside from the syntax and RNG differences described earlier, the two implementations look basically the same!
However, one interesting difference to note here is how causal masks are handled. In PyTorch, it’s common to register the mask as a buffer at initialization:
self.register_buffer(
"causal_mask",
torch.tril(torch.ones(max_seq_len, max_seq_len, device=device)),
persistent=False,
)Unlike parameters, buffers are not learnable and not tracked by the optimizer. However, they are automatically moved with the model during device placement.
In JAX, we can just assign it directly:
self.causal_mask = jnp.tril(jnp.ones((max_seq_len, max_seq_len)))This is because device placement is automatic in JAX, so there’s no need for a special registration mechanism to ensure the mask follows the model to the correct device. As a bonus, since jax.jit treats any closed-over array as a constant in the compiled graph, the mask is never recomputed across calls.
The other components of our language model (SwiGLU, RMSNorm, Embedding, LM Head) follow much of the same design. We recommend taking a look at the full model implementation in JAX, which you can find here.
Implementing the Training Loop
Now that we have our model implemented in JAX, let’s take a look at how we can train it!
There are a few major differences in the JAX training loop compared to PyTorch. We’ll discuss each one and what it might reveal about the JAX paradigm.
The Training Step
In PyTorch, our train step looks something like this:
logits = model(inputs)
B, S, V = logits.shape
loss = model_module.cross_entropy_loss(logits.reshape(B * S, V), targets.reshape(B * S))
optimizer.zero_grad()
loss.backward()
optimizer.step()Meanwhile, in JAX, it looks something like:
loss, grad_state = train_step(model, optimizer, inputs, targets)Where the train_step function looks something like:
@nnx.jit
def train_step(model: nnx.Module, optimizer: nnx.Optimizer, inputs: Array, targets: Array):
def loss_fn(model):
logits = model(inputs)
B, S, V = logits.shape
return cross_entropy_loss(logits.reshape(B * S, V), targets.reshape(B * S))
loss, grads = nnx.value_and_grad(loss_fn)(model)
optimizer.update(model, grads)
return loss, gradsTrain Step Function: In PyTorch, we don’t need to wrap our training step into a function. Most of our operations operate in place. However, we need to create a train_step function in JAX so that we can wrap it with nnx.jit.
Gradient Handling: In PyTorch, gradients are implicit, since loss.backward() accumulates gradients into each parameter’s .grad field, and optimizer.step() reads those .grad buffers and updates parameters in place. As a result, our gradient states are effectively distributed across {p.grad for p in model.parameters()}, instead of being unified in a single gradient object grad like in JAX. Gradient clipping in PyTorch follows the same philosophy by scaling the .grad buffers in place.
On the other hand, gradients are treated as explicit values in JAX. To get our grad object, we use nnx.value_and_grad, which is a transformation that turns the loss_fn into a function that also returns gradients. Since JAX transforms act as higher order functions, we need to define a loss function and pass it in as input to nnx.value_and_grad instead of just calling loss.backward() like we do in PyTorch.
Now that we have the basic training step, let’s add gradient clipping and a learning rate schedule.
Gradient Clipping and LR Schedule
If we wanted to add gradient clipping and a learning rate schedule in PyTorch, they would be called separately from our optimizer. In JAX, they would be processed by the optimizer internally.
PyTorch Here’s what it would look like in PyTorch:
def build_lr_scheduler(
optimizer: torch.optim.Optimizer,
min_learning_rate: float,
warmup_iters: int,
cosine_annealing_iters: int,
warmup_start_factor: float = 0.1,
) -> torch.optim.lr_scheduler.LRScheduler:
cosine_decay_iters = cosine_annealing_iters - warmup_iters
warmup_scheduler = torch.optim.lr_scheduler.LinearLR(
optimizer,
start_factor=warmup_start_factor,
end_factor=1.0,
total_iters=warmup_iters,
)
cosine_scheduler = torch.optim.lr_scheduler.CosineAnnealingLR(
optimizer,
T_max=cosine_decay_iters,
eta_min=min_learning_rate,
)
scheduler: torch.optim.lr_scheduler.LRScheduler = torch.optim.lr_scheduler.SequentialLR(
optimizer,
schedulers=[warmup_scheduler, cosine_scheduler],
milestones=[warmup_iters],
)
return schedulerand the new train_step would look something like this:
logits = model(inputs)
B, S, V = logits.shape
loss = model_module.cross_entropy_loss(logits.reshape(B * S, V), targets.reshape(B * S))
optimizer.zero_grad()
loss.backward()
if gradient_clip > 0: # gradient clipping
torch.nn.utils.clip_grad_norm_(model.parameters(), float(gradient_clip))
optimizer.step()
if scheduler is not None: # scheduler handling
scheduler.step()In the function build_lr_scheduler, we create a scheduler object that we chains together a linear warmup schedule with a cosine annealing schedule. To use this, we call scheduler.step(), which computes new LR values and mutates the optimizer’s param_groups’ lr attributes in place.
JAX In contrast, gradient clipping and the learning rate scheduler are used as inputs during optimizer construction in JAX. Our optimizer construction would look something like:
lr_schedule = optax.warmup_cosine_decay_schedule(
init_value=0.0,
peak_value=a_max,
warmup_steps=warmup_iters,
decay_steps=decay_iters,
end_value=a_min
)
def build_optimizer_transform(
optimizer_config: dict,
gradient_clip: float,
lr_schedule: optax.Schedule,
) -> optax.GradientTransformation:
transforms = []
if gradient_clip > 0:
transforms.append(optax.clip_by_global_norm(float(gradient_clip)))
transforms.append(
adamw(
lr=lr_schedule,
betas=tuple(optimizer_config["betas"]),
weight_decay=float(optimizer_config["weight_decay"]),
eps=float(optimizer_config["eps"]),
)
)
return optax.chain(*transforms)
optimizer = nnx.Optimizer(
model,
build_optimizer_transform(
optimizer_config,
gradient_clip=gradient_clip,
lr_schedule=lr_schedule,
),
wrt=nnx.Param,
)In JAX, the optimizer is treated as a series of Optax transformations. We can first construct our pipeline by adding gradient clipping followed by AdamW. Note that we pass in our learning rate schedule lr_schedule as an argument to AdamW during initialization. Then, we can create an nnx.Optimizer using our model, the pipeline we just created, and the wrt argument that specifies what to optimize.
Finally, we can call this without changing our code by using optimizer.update(model, grads), which will handle the gradient clipping and LR updates internally. Note that unlike PyTorch, we don’t need to call scheduler.step() or clip_grad_norm_() explicitly in each step.
You can find the full training script here.
Intro to Distributed Training
So far, we have assumed that both the model and data batch fit on a single device. However, once the model state or activations become too large, or once we want more throughput than a single device can provide, we need to spread (a.k.a shard) the work across multiple GPUs/TPUs. Doing so can give us more total memory and compute, but it also introduces a new bottleneck: communication. As a result, a core problem in distributed training is deciding which tensors stay local to each device, which tensors are sharded, and which tensors should be communicated between devices.
Different parallelism strategies answer this question in different ways. In this section, we will focus on 4 common strategies: Data Parallelism (DP), Fully Sharded Data Parallelism (FSDP), Tensor Parallelism (TP), and mixed FSDP+TP. A useful way to think about them is:
- DP splits the data batch
- FSDP splits the persistent model state
- TP splits the hidden-state computation inside a layer
- FSDP+TP does both at once
Don’t worry if this doesn’t make much sense yet! The goal of this section is to build some intuition for these strategies, and in the next section we will show you how to implement them in JAX.
Note
This section is not meant as a complete guide to distributed training. We recommend starting with the How to Scale Your Model book for a more complete guide.
Notation
We will denote variables in the form $\text{VarName}[d_1, d_2, \ldots]$ giving its name and shape, respectively. To keep the notation simple, we will approximate a Transformer block as a pair of linear layers:
\[\text{In}[B, D] \cdot W_{1}[D, F] \cdot W_{2}[F, D] \to \text{Out}[B, D]\]Why can we use this approximation?
A Transformer block contains two main sub-blocks: multi-head attention (MHA) and the feed-forward network (FFN). Both include large projection matrices that map from the model dimension $D$ into some intermediate space and then back to $D$.
For the FFN, this is exactly the $D \to F \to D$ pattern. For MHA, the input is first projected into Q/K/V representations, attention is applied, and the result is then projected back into $D$ by the output weights $W_O$.
We simplify both sub-blocks to a pair of linear layers since we care mostly about how we shard the large projection matrices and the communication they induce. The operations in-between (activations, attention, layernorms) are typically performed locally on each device under the sharding strategies we discuss. As a result, they typically don’t introduce additional communication.
In many common LLM settings, the FFN also accounts for a large fraction of the block FLOPs, which makes this simplification especially useful.
One important note here is that $\text{In}[B, D]$ represents the input activations, not necessarily of the raw input batch itself. In language modeling, we typically have a tokenized input batch of shape $[B, T]$ where $T$ is the number of tokens per sample. This input batch would then be fed through our embedding table to get our activations $[B, T, D]$, which is what we refer to here as $\text{In}$. Following common practice, we abuse our notation a bit to roll the $B$ and $T$ dimensions into one $B$ that represents the total number of tokens in a batch.
For sharding, we will use subscripts like $B_X$ and $D_Y$ to denote sharding over device-mesh axes $X$ and $Y$, respectively. For example, $B_X$ means the batch dimension is split across the $X$-axis of the device mesh, while $D_Y$ means that the hidden dimension is split across the $Y$-axis. We’ll introduce the device mesh in the next section.
In our illustrations, we will mostly show weight gradients and omit input gradients when they aren’t the main point of the conversation.
Before we dive into parallelism schemes, let’s take a look at some important preliminaries: the device mesh, and communication collectives.
The Device Mesh
To talk about sharding, we need to have a way to refer to our devices. The standard abstraction is a device mesh, which you can imagine as arranging all of our devices into a matrix. For example, if we had 8 devices, we might arrange them as a $4 \times 2$ mesh, a $8 \times 1$ mesh, a $2 \times 4$ mesh, and so on. Each dimension of the mesh is called a device axis. In this section, we’ll refer to these axes abstractly as $X$ and $Y$. Later, in JAX, we will assign them names like "data" "tensor".
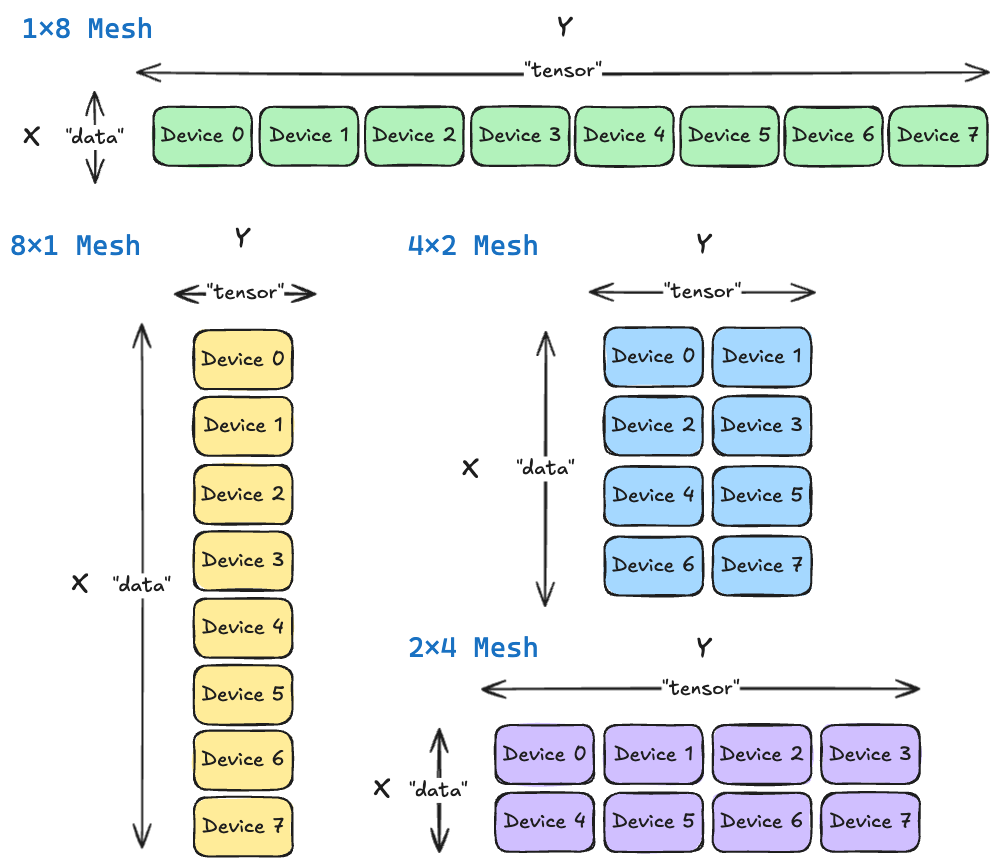
Communication Collectives
When we talk about parallelism, there are also 3 communication operations that will pop up repeatedly. These are the All-Gather, Reduce-Scatter, and All-Reduce.
All-Gather
During an all-gather operation, each node sends its shard of data to all other nodes, so each device can reconstruct the whole. When we have shards of a tensor on different devices, we can use an all-gather to build a copy of the full tensor on each device.

Reduce-Scatter
Each device computes the result on their own shard of data, but a “reduce” operation is needed to compute the overall result incorporating all shards. Then, the final result must be re-sharded and distributed back to the corresponding device—this is the “scatter” operation. Reduce-scatter combines these operations into a single step.
It does this using a ring structure, illustrated below. Devices are organized in a loop and each device passes a segment to its neighbor and receives one from the other neighbor, repeating for $N-1$ steps. By the end, each device is holding their distinct, fully reduced segment of the data.

All-Reduce
An all-reduce combines partial results across devices (often a sum) and leaves every device with a copy of the full result. Equivalently, you can think of it as a reduce-scatter (combine and distribute one shard per device) followed by an all-gather (collect all shards so every device has the complete result).
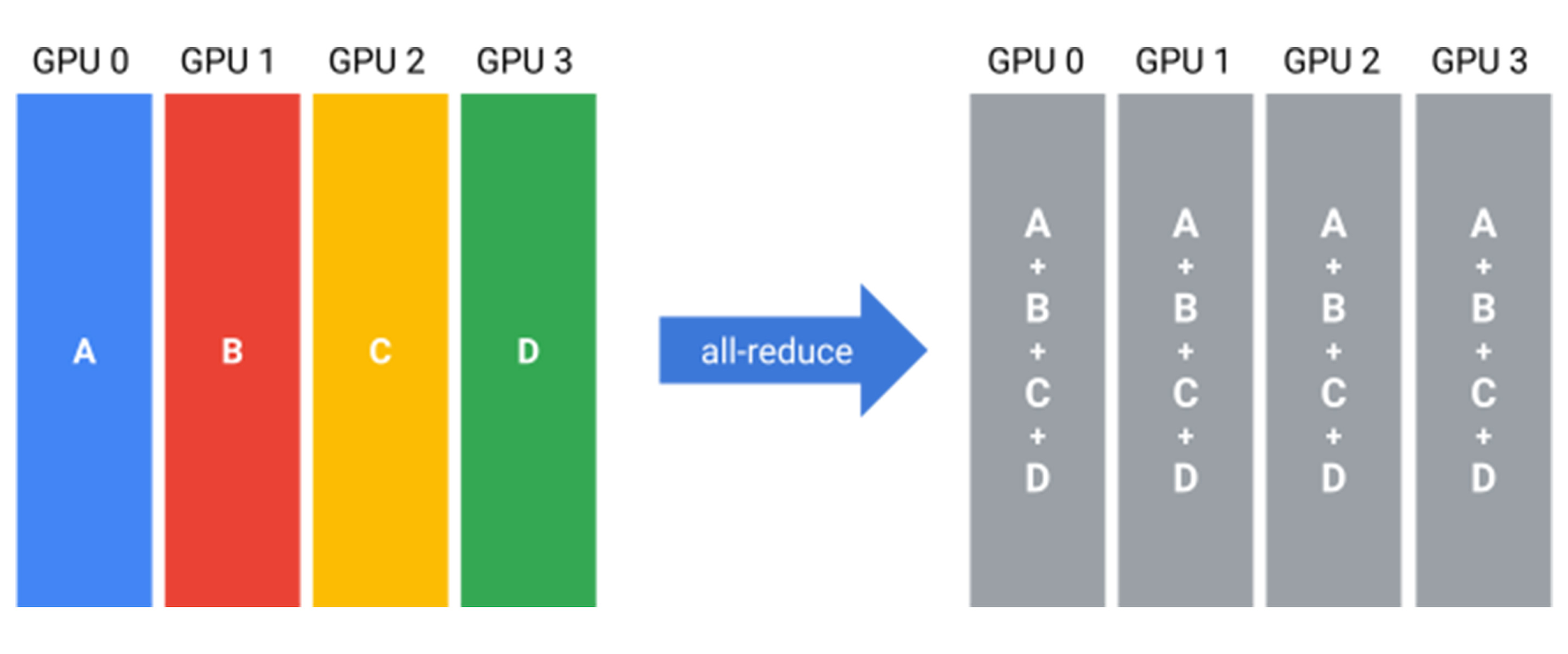
The beauty of JAX is that we never need to implement or code these operations ourself. This is all handled implicitly once we define the device mesh. Now, with preliminaries out of the way, let’s talk about training!
Standard Training
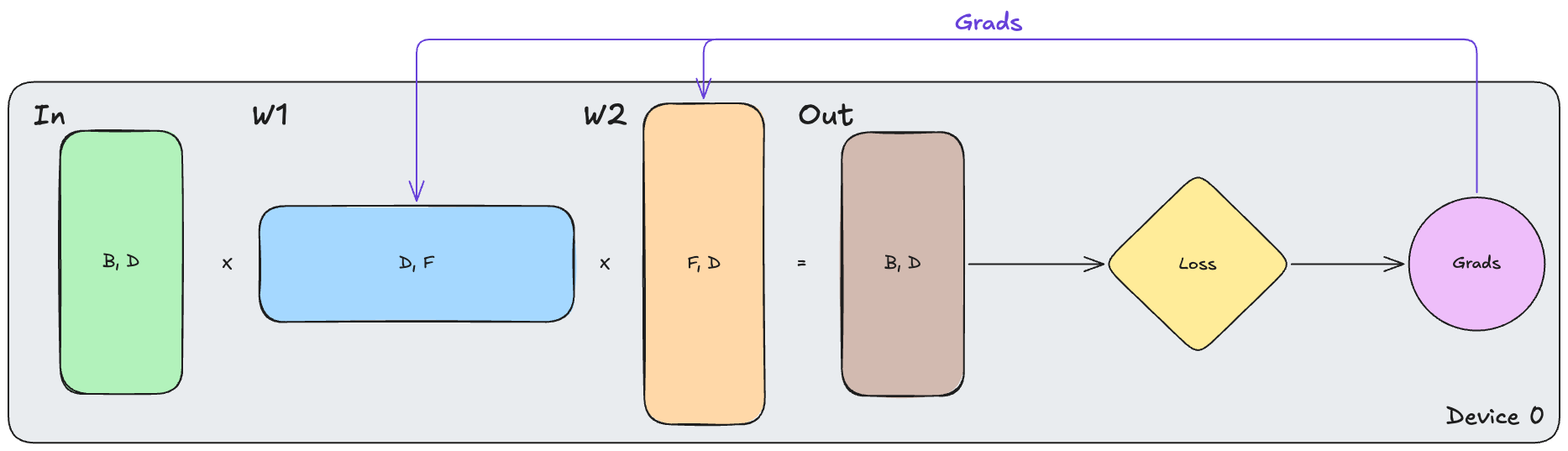
As a starting point, let’s take a look at the non-distributed case. We can represent this with the following:
\[\text{In}[B, D] \cdot_{D} W_{1}[D, F] \cdot_{F} W_{2}[F, D] \to \text{Out}[B, D]\]On a single device, the full input, weights, activations, gradients, and optimizer state all live together. The forward pass computes $\text{Out}$, the backward pass computes gradients $\text{d}W_1$, $\text{d}W_2$, $\text{d}\text{In}$, and the optimizer uses the gradients to update the weights $W_1, W_2$. This is the simplest case, and it gives us a reference point for the distributed strategies below.
Data Parallel (DP)
Data parallelism is the simplest distributed strategy. We shard the input across the batch dimension and keep a full copy of the weights and optimizer state on every device.
\[\text{In}[B_X, D] \cdot W_{1}[D, F] \cdot W_{2}[F,D] \rightarrow \text{Out}[B_X, D]\]Each device therefore runs the same forward and backward computation, but on a different slice of the batch. Because every device has the full weights locally, the forward pass requires no communication.
However, after the backward pass, each device only has a local contribution to the weight gradients from its own slice of the batch. These are displayed as Grads1 and Grads2 in the diagram below illustrating DP on a 2x1 mesh. To recover the gradient for the full batch, we need to all-reduce those gradients across devices. Each device then receives the same summed gradient Grads and applies the same update to the same weights, which ensures that our weights are always in sync across devices.
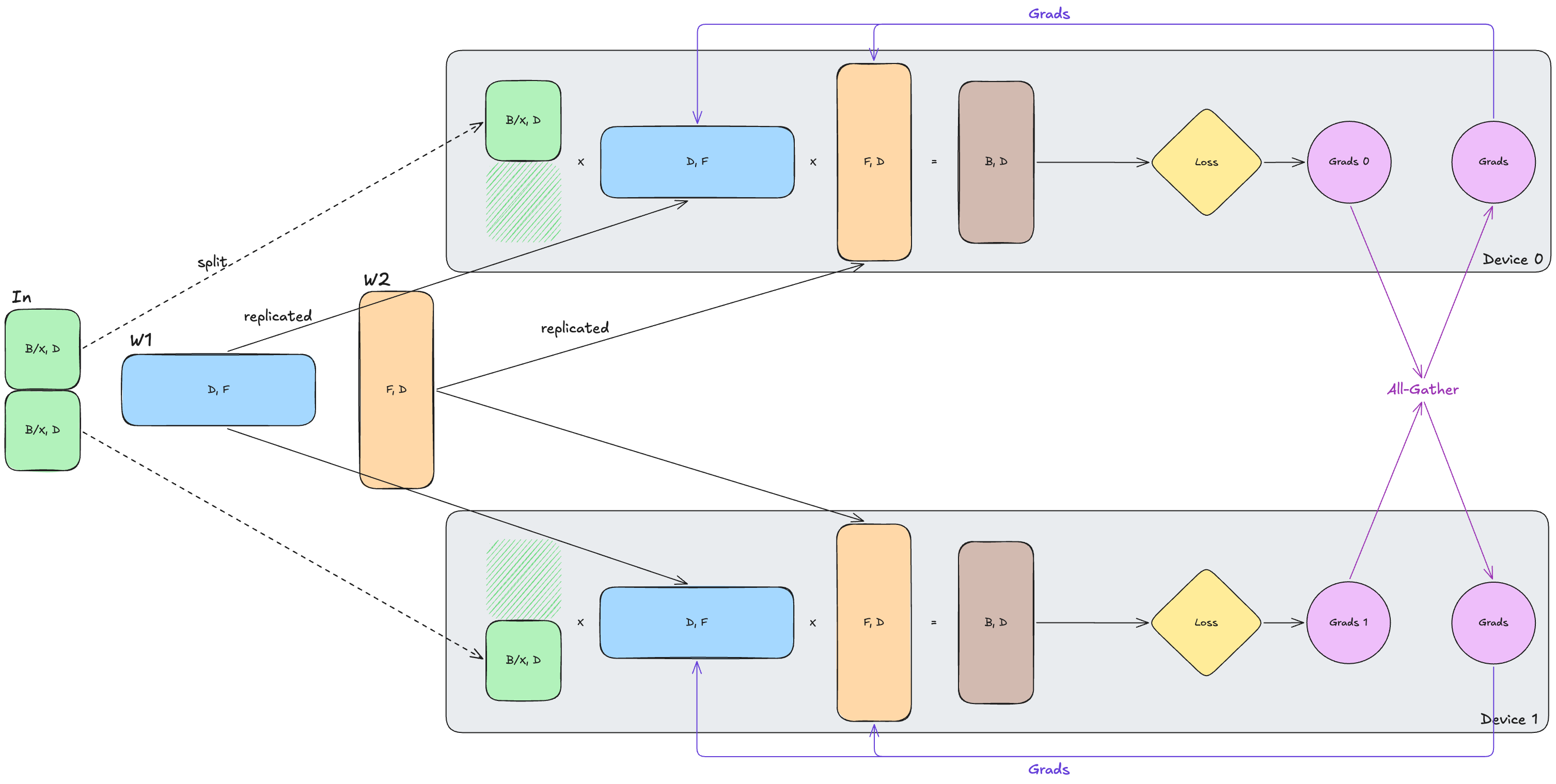
To summarize, the main ideas of DP are:
- Cheap forward pass
- Replicated model state
- Synced gradients in the backward pass
Fully-Sharded Data Parallel (FSDP)
You might notice a disadvantage of DP is that each device must hold a complete copy of the weights, gradients, and optimizer state. FSDP tries to solve this by sharding the model state across devices as well:
\[\text{In}[B_X, D] \cdot W_{1}[D_{X}, F] \cdot W_{2}[F,D_{X}] \rightarrow \text{Out}[B_X, D]\]Now, each device only permanently stores its own shard of the weights and optimizer state. The tradeoff here is that a device usually can’t perform the forward pass matmul using only its local weight shard, since our weights are sharded across the contracting dimension $D$. As a result, we need to all-gather the parameter shards so that each device can materialize the full weight matrix before the layer runs. After this forward pass computation, the temporary full copy can be discarded.
In the backward pass, each device will only have the gradients of the local slice of the input, similar to DP. However, instead of all-reducing the full gradient like DP, we can reduce-scatter the gradient so that each device keeps only the gradient shard corresponding to the parameters it owns. The key idea is that the FSDP lowers the persistent per-device memory footprint, even though each device still temporarily materializes the full layer weights when the layer is executing.
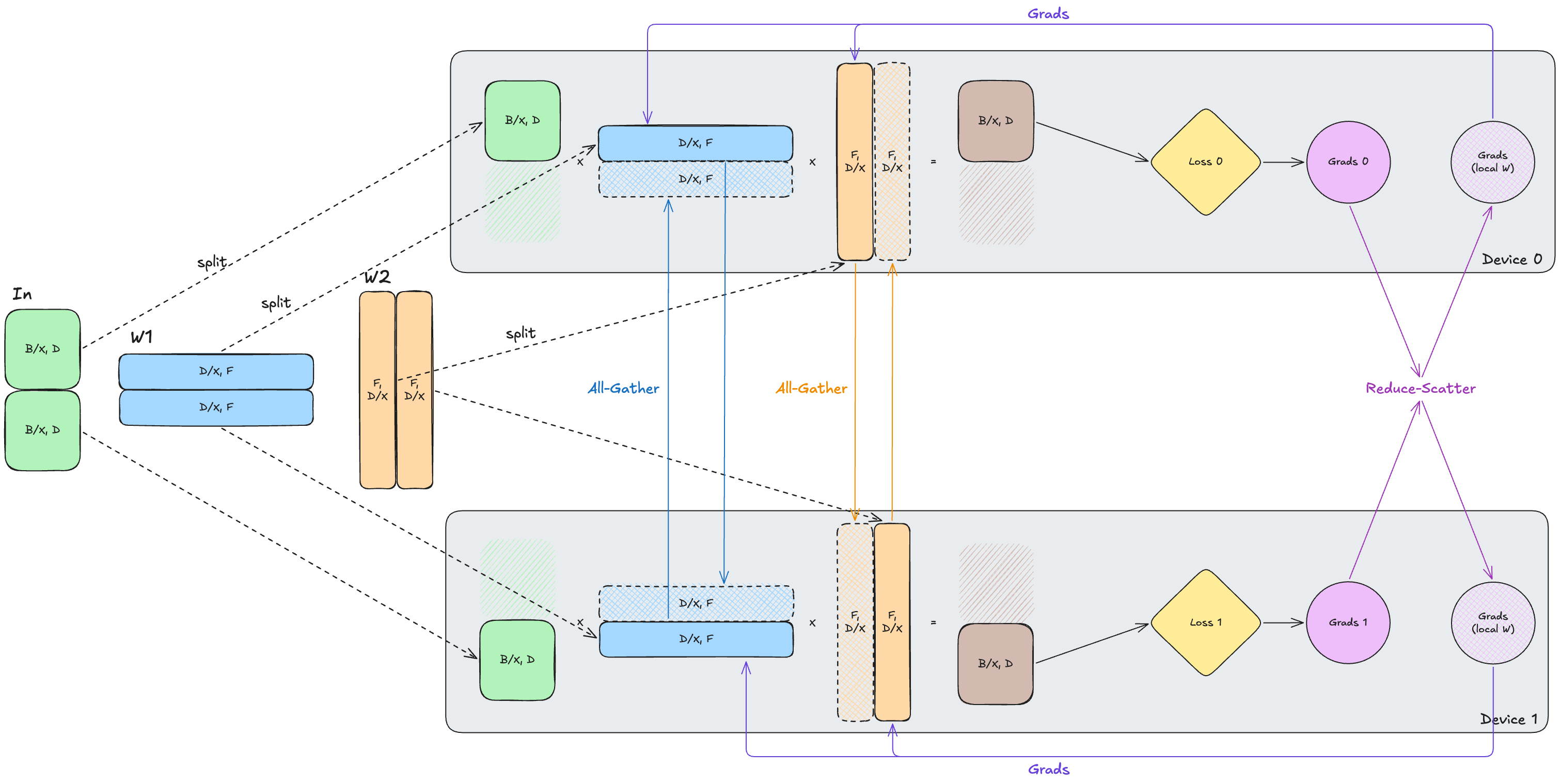
In this illustration, the crosshatched/dashed weight shards are received from the all-gather and discarded after use.
Tensor Parallel (TP)
One important observation is that FSDP mostly reduces persistent parameter memory but not necessarily the peak per-device memory during the forward pass. As a result, if our model or sequence length is so large that our activations don’t fit in a single forward pass, FSDP won’t help us much. Tensor parallelism (TP) takes a different approach by sharding the computation itself, rather than only sharding stored parameters. Instead of moving weights across devices, we shard the weights and distribute the computation of a single layer across devices. Here is how we can represent TP:
\[\text{In}[B, D] \cdot_D W_{1}[D, F_{Y}] \cdot_{F} W_{2}[F_{Y},D] \rightarrow \text{Out}[B, D]\]Here, we shard along the feature dimension (denoted by $Y$), so each device holds only a slice of the intermediate activations and weights. In this setup, each device computes only its local contribution to the output. Importantly, since each device can compute the gradient for its local weight shard directly, no communication is required for weight gradients.
However, communication is still required elsewhere. After the forward pass, each device holds only a partial output from its local weight shards, so we must all-reduce our output across devices to obtain the full output needed for the loss. In the backward pass, the same issue appears for the input gradients: each device computes only a partial contribution to $\text{dIn}$, so we again need an all-reduce to combine these contributions (although input gradients are not shown in the diagram for simplicity).
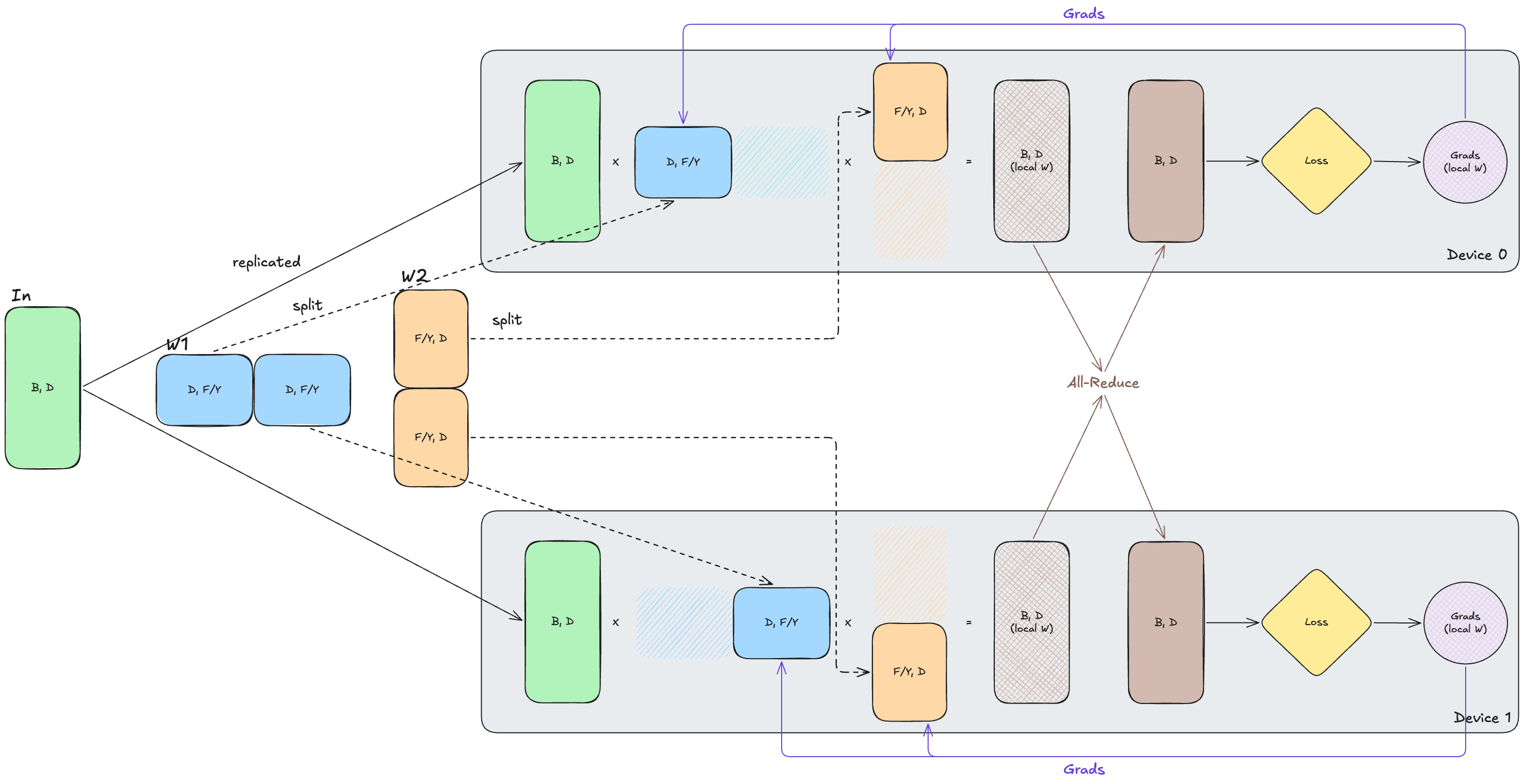
FSDP + TP
Can we take FSDP and TP a step further? Turns out we can actually combine them by assigning each to different device axes! Along the $X$ axis, we use FSDP to shard model state, while along the $Y$ axis, we use TP to shard the hidden-state computation:
\[\text{In}[B_{X}, D_{Y}] \cdot_D W_{1}[D_{X}, F_{Y}] \cdot_{F} W_{2}[F_{Y},D_{X}] \rightarrow \text{Out}[B_{X}, D_{Y}]\]This combination is useful because the two strategies help each other. FSDP shards the batch across the $X$-axis, which reduces the amount of activation data that TP must move. TP shards the weights across $Y$-axis, which reduces the amount of parameter data that FSDP must move. The tradeoff is that we now pay both activation communication and weight communication, so the implementation is more complex and communication costs are higher as well. For a discussion on how to balance FSDP and TP, and when to use it, we recommend reading the related section in the How to Scale Your Model book.
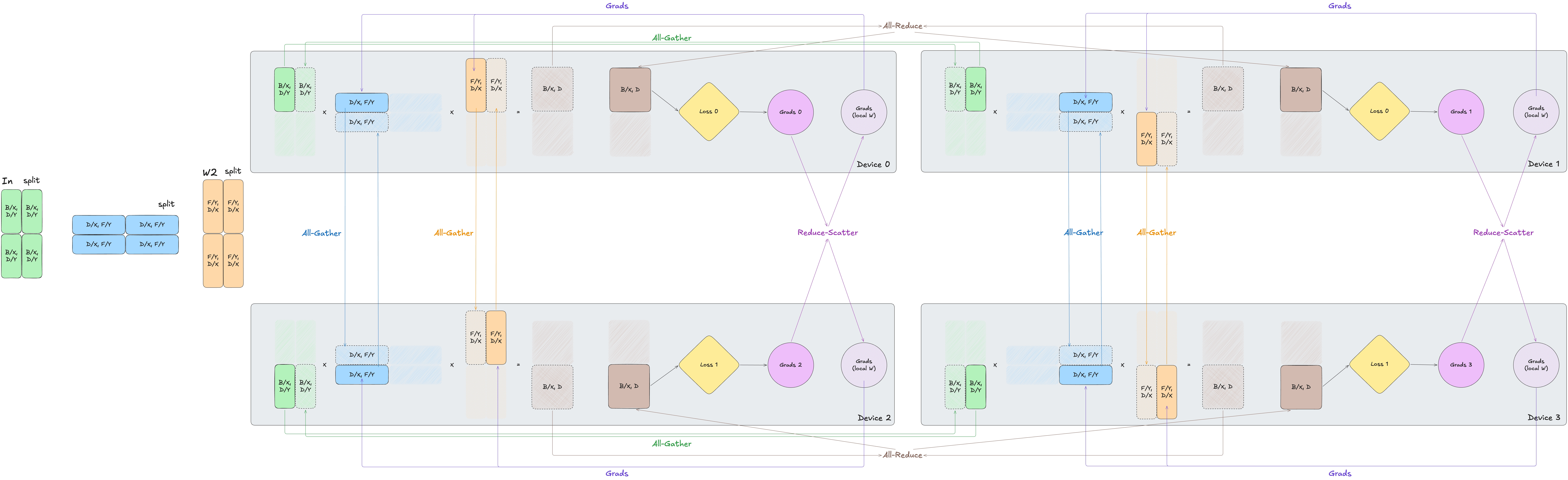
This image might be a bit hard to read due to the size, but you can find the full-size image here.
In this example, we expand the number of devices we use to 4, since we need to use at least 2 devices to perform FSDP and another 2 to perform TP. We also omit the arrows showing the initial splitting of weights, but each device will hold a shard of both the input and the weights, corresponding to its position in the mesh.
Before the matmul, TP all-gathers the inputs across $Y$ to get the full data dimension $D$. Then, we can use FSDP to gather our weight matrices across the $X$ axis. Then, we perform a TP-like All-Reduce for our output shards across the $Y$ axis, then finally reduce scatter our gradients across $X$ in an FSDP-like way. One thing to note here is that since we perform an all-reduce of the partial $\text{Out}[B_X, D]$ tensors across the $Y$-axis, each row will actually have the same loss.
Implementing Distributed Training
Now that we have a conceptual picture of distributed training, implementing it in JAX is mostly about expressing the same layouts explicitly. We define a device mesh, specify which tensor dimensions are sharded over which mesh axes, and let JAX/XLA insert the necessary collective communications.
How does this compare to PyTorch?
In PyTorch’s distributed training stack, the programming model is similar (we create a device mesh and specify how tensors are sharded) but the underlying execution is different, as PyTorch relies on explicit collective wrappers. For example, FSDP registers pre-forward/post-forward hooks that call all-gather and reduce-scatter at precise points, and Tensor Parallel inserts all-reduce or reduce-scatter calls around colwise/rowwise parallel layers. In JAX, we don’t write these collectives ourselves since XLA takes care of this automatically. This means that switching from something like DP to FSDP+TP in JAX just requires us to change our partition specs and mesh shape, whereas in PyTorch it would often require swapping wrapper classes and restructuring the training loop.
In this section, we will go over:
- Initializing our device mesh
- Sharding annotations
- How to use sharding annotations to initialize the model
- How to define our sharding annotations for each parallelism mode
Let’s begin!
Initializing our Device Mesh
In the previous section, we referred to our mesh axes as $X$ and $Y$. We can now give these axes names, which are conventionally named "data" and "tensor". The "data" axis is used for DP/FSDP-style sharding, while the "tensor" axis is used for TP-style sharding.
We can define our device mesh as following:
from jax.sharding import Mesh
def create_mesh(mesh_shape: list[int], mesh_axis_names: list[str]) -> Mesh:
auto_mesh = jax.make_mesh(tuple(mesh_shape), tuple(mesh_axis_names))
return auto_mesh
mesh = create_mesh((4, 2), ("data", "tensor"))Here, we use the jax.make_mesh() function to create our Mesh object. We pass in (4, 2) as our mesh shape to create a 4x2 device mesh, with the "data" being the X device axis and "tensor" being the Y device axis.
Attaching Sharding Annotations to our Model
Now that we have our device mesh, our next step is to specify how we want our data and model to be sharded on this mesh. We can use the PartitionSpec class to do this, which is a tuple abstraction that defines our sharding annotations.
For example, after importing PartitionSpec using from jax.sharding import PartitionSpec as P, as is customary, we can define:
-
P("data", None), which means that we shard the 1st array dimension across the “data” axis and leave the second dimension unsharded -
P(None, "tensor")means that we shard the 2nd array dimension across the “tensor” axis and leave the first dimension unsharded -
P(None, None)means that we do not shard either array dimension. Essentially, this means that the tensor is replicated across devices. To make this concrete, if I had a device mesh of shape $4 \times 2$ and a tensor $X[32, 8]$, thenP("data", None)will result in a $X[32/4, 8] = X[8, 8]$ tensor that is replicated across the $Y$ axis. If I useP("data", "tensor")instead, each device would have a $X[32/4, 8/2] = X[8, 4]$ slice of the original tensor.
The diagram below illustrates several sharding configurations for our 4×2 mesh. Note that the notation Device A,B,C indicates that slice of the matrix is replicated across devices $A$, $B$, and $C$.
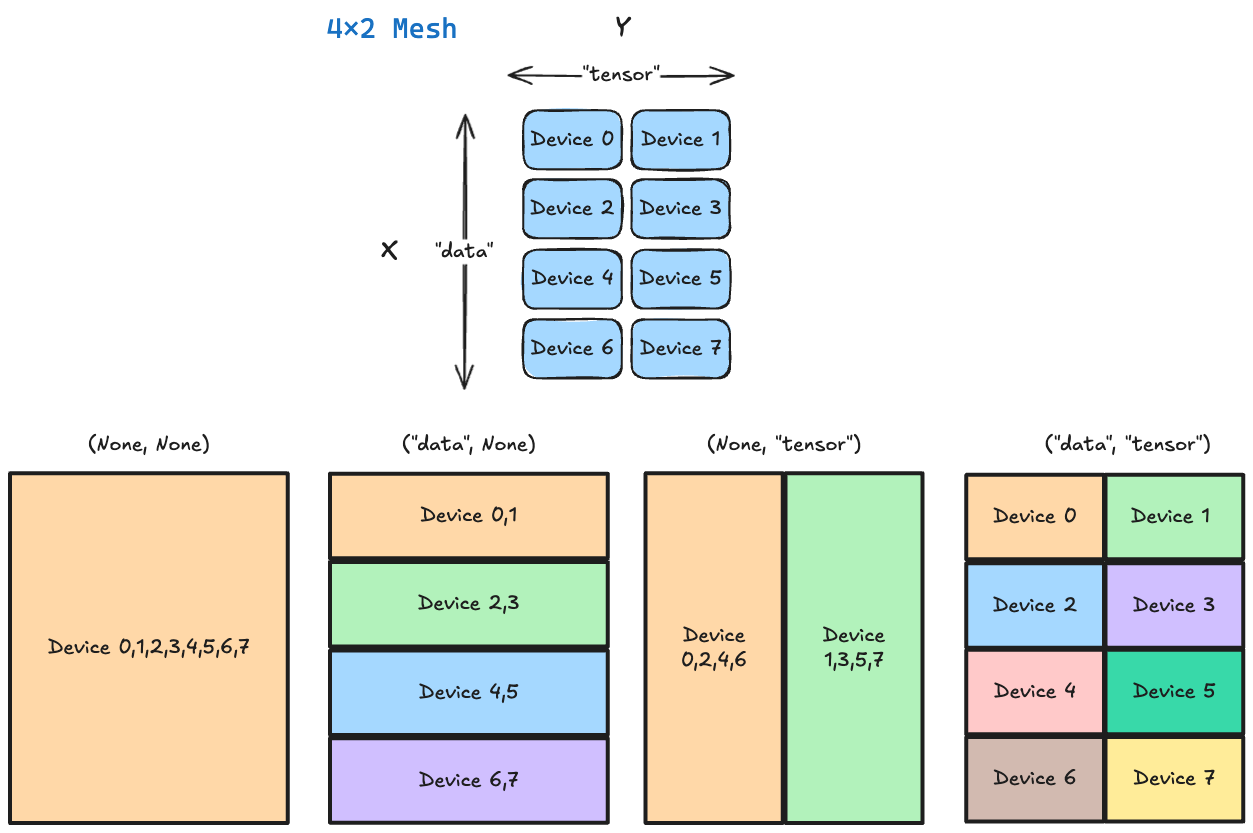
For now, you don’t need to worry about what specific PartitionSpec to use, as it depends on what parallelism strategy we choose.
Initializing our Model with Sharding
The next step is to attach these sharding annotations to the tensors that we want to shard. In NNX, the easiest way to do this is to wrap the initializer with nnx.with_partitioning(init_fn, sharding), which will attach sharding annotations to the variable created with init_fn. Typically, these sharding annotations should be passed in at model initialization so that JAX knows how to distribute tensors across devices.
Here’s what it might look like in code:
class Linear(nnx.Module):
def __init__(self, rngs: nnx.Rngs, in_features: int, out_features: int,
sharding: Sharding | None = None, dtype: jnp.dtype = jnp.float32,
):
super().__init__()
std = (2 / (in_features + out_features)) ** 0.5
# We use `nnx.with_partitioning` here!
init_fn = nnx.with_partitioning(
nnx.initializers.truncated_normal(stddev=std, lower=-3.0 * std, upper=3.0 * std), sharding
)
weights_data = init_fn(rngs.params(), (in_features, out_features), dtype)
self.weights = nnx.Param(weights_data)
def __call__(self, x: jnp.ndarray) -> jnp.ndarray:
return jnp.einsum("...i,io->...o", x, self.weights.get_value())The only differences from a standard Linear layer are that:
- We wrap the original
nnx.initializers.truncated_normal()call with thennx.with_partitioning()function - We pass in the
shardingas input The process is the same for other components of the model, all we need to do is just wrap the initializer withnnx.with_partitioning.
Now that we’ve annotated each component of our model with nnx.with_partitioning, we need to actually initialize our model so that our tensors are created on the right devices. The cleanest initialization pattern is to create the model inside a mesh context manager and JIT-compile the initialization for performance.
Here’s what it might look like in code:
import optax
@jax.jit
def init_model_and_optimizer(rngs: nnx.Rngs, model_config, optimizer_config, sharding_config):
model = TransformerLM(
rngs=rngs,
model_config=model_config,
sharding_config=sharding_config,
)
optimizer = nnx.Optimizer(
model,
build_optimizer_transform(
optimizer_config,
gradient_clip=optimizer_config["gradient_clip"],
lr_schedule=optimizer_config["lr_schedule"],
),
wrt=nnx.Param,
)
return model, optimizer
with jax.set_mesh(mesh):
model, optimizer = init_model_and_optimizer(
nnx.Rngs(0),
model_config,
optimizer_config,
sharding_config,
)Sharding our Data
Beyond the model itself, we also need to shard the input data. The approach depends on the parallelism strategy: in TP, each device needs the full batch, while in DP, FSDP, and combined FSDP+TP we split the batch across the data axis so each device processes a different slice.
We can capture this logic with a simple helper:
def get_batch_sharding_for_mode(mode: str) -> Sharding:
if mode == "tp":
return (None, None) # replicate across all devices
if mode in ("dp", "fsdp", "fsdp_tp"):
return ("data", None) # shard batch over data axis, replicate over tensor axis
raise ValueError("invalid mode!")
batch_partition_spec = P(*model_module.get_batch_sharding_for_mode(sharding_mode))Why is FSDP+TP not (“data”, “tensor”)?
If you are paying attention, you might notice that in the FSDP+TP formula, $\text{In}$ is sharded as $\text{In}[B_X, D_Y]$. This would equate to ("data", "tensor"), but we instead have ("data", None) here. Why?
Recall that the raw batches we load from the dataset are not the same tensors as the hidden activations used in the simplified formulas above.
- The dataloader returns token IDs and targets, which usually have shape $[B, T]$.
- The notation $\text{In}[B_X, D_Y]$ in the previous section refers to the input activations after embedding into the model dimension $D$ to get $[B, T, D]$.
Because raw token batches don’t have a hidden dimension $D$ yet, even in combined FSDP+TP we only shard them over the "data" axis and replicate them over the "tensor" axis. The tensor-axis sharding appears later, once the model has produced hidden states.
Then, at data loading time, we use jax.device_put to place the batch according to this spec:
def get_batch_from_memmap(
rngs: nnx.Rngs,
dataset: DatasetLike,
batch_size: int,
context_length: int,
mesh: Mesh | None = None,
batch_partition_spec: P | None = None,
) -> tuple[Array, Array]:
inputs, targets = model_module.get_batch(
rngs,
dataset.data,
batch_size,
context_length,
)
# Place batch on devices according to the partition spec
if mesh is not None:
with jax.set_mesh(mesh):
inputs = jax.device_put(inputs, batch_partition_spec)
targets = jax.device_put(targets, batch_partition_spec)
return inputs, targetsDefining Our Sharding
Before discussing each parallelism strategy separately, it’d be helpful to first identify the major tensors in the model and the sharding pattern each one will use. Our model contains the following components:
| Component | Shape |
|---|---|
| $\text{Input}$ | $[B, D]$ |
| $W_{Q}, W_{K}, W_{V}$ | $[D, N*H]$ |
| $W_{O}$ | $[N*H, D]$ |
| $W_{1}, W_{3}$ | $[D, F]$ |
| $W_{2}$ | $[F, D]$ |
| $\text{RMSNorm}$ | $[D]$ |
| $\text{Embedding}$ | $[V, D]$ |
| $\text{LM Head}$ | $[D, V]$ |
Rather than assigning every tensor a completely separate sharding rule, we can group tensors by the role they play in the computation.
In particular, many weight matrices fall into one of two common patterns:
- Projection out of the model dimension $D$ into some larger or different dimension. This includes $W_{Q}, W_{K}, W_{V}$ and $W_{1}, W_{3}$, which have shape $[D, *]$. We can refer to this pattern as
dense_in - Projection back into the model dimension $D$ from some intermediate dimension. This includes $W_{O}$ and $W_{2}$. We will refer to this pattern as
dense_out. This naming is useful because these matrices often share the same partitioning logic under a given parallelism strategy, even if their exact shapes differ.
What about Embedding and LM Head?
You might notice that the Embedding and LM Head (the unembedding matrix) are not grouped into dense_in and dense_out. This is because they often play a special role because they involve the vocabulary dimension $V$ instead of hidden dimensions like $D$, $F$, or $N*H$. As a result, they often need different sharding rules since the embedding performs lookup over the vocabulary rows, while the LM head produces vocab logits, which may be sharded across vocab and may require special handling for softmax and cross-entropy. Finally, these 2 layers are often also weight-tied in many language models, which adds an extra layout constraint. For these reasons, it’s cleaner to keep them as separate sharding categories.
With this grouping, we can assign each component a sharding label:
| Component | Shape | Sharding Name |
|---|---|---|
| $\text{Input}$ | $[B, D]$ | batch |
| $W_{Q}, W_{K}, W_{V}$ | $[D, N*H]$ | dense_in |
| $W_{O}$ | $[N*H, D]$ | dense_out |
| $W_{1}, W_{3}$ | $[D, F]$ | dense_in |
| $W_{2}$ | $[F, D]$ | dense_out |
| $\text{RMSNorm}$ | $[D]$ | norm |
| $\text{Embedding}$ | $[V, D]$ | embedding |
| $\text{LM Head}$ | $[D, V]$ | lm_head |
These names only serve as categories for now. In the next step, for each parallelism strategy, we will map each one to an actual sharding specification: the tuple of strings describing the 2D partition over the tensor’s axes. For 1D tensors such as RMSNorm, the corresponding specification will be 1D rather than 2D.
Now, let’s take a look at how we can define what each of these sharding names should be under data parallelism, FSDP, tensor parallelism, and FSDP+TP!
Shardings for Each Strategy
DP
In Data Parallel, our basic formula looks something like the following:
\[\text{In}[B_X, D] \cdot_D W_{1}[D, F] \cdot_F W_{2}[F, D] \rightarrow \text{Out}[B_X, D]\]We want to split our batches across the $X$, or “data” axis, and replicate our weights otherwise.
| Component | Shape | Sharding Name | DP | Resulting Shape (per device) |
|---|---|---|---|---|
| $\text{Input}$ | $[B, D]$ | batch | ("data", None) | $\left[ \frac{B}{X}, D \right]$ |
| $W_{Q}, W_{K}, W_{V}$ | $[D, N*H]$ | dense_in | (None, None) | $[D, N*H]$ |
| $W_{O}$ | $[N*H, D]$ | dense_out | (None, None) | $[N*H, D]$ |
| $W_{1}, W_{3}$ | $[D, F]$ | dense_in | (None, None) | $[D, F]$ |
| $W_{2}$ | $[F, D]$ | dense_out | (None, None) | $[F, D]$ |
| $\text{RMSNorm}$ | $[D]$ | norm | (None) | $[D]$ |
| $\text{Embedding}$ | $[V, D]$ | embedding | (None, None) | $[V, D]$ |
| $\text{LM Head}$ | $[D, V]$ | lm_head | (None, None) | $[D, V]$ |
Remember that None replicates the dimension of the tensor across that axis!
Here’s what it might look like in code:
def get_sharding_config_for_mode(mode: str) -> ShardingConfig:
if mode == "dp":
return ShardingConfig(
dense_in=(None, None),
dense_out=(None, None),
embedding=(None, None),
lm_head=(None, None),
norm=(None,),
)
...And that’s all we need to do! Simple, right?
FSDP
In FSDP, we have the following:
\[\text{In}[B_X, D] \cdot_D W_{1}[D_X, F] \cdot_F W_{2}[F, D_X] \rightarrow \text{Out}[B_X, D]\]We want to shard both our input activations and weights along the data axis.
| Component | Shape | Sharding Name | FSDP | Resulting Shape (per device) |
|---|---|---|---|---|
| $\text{Input}$ | $[B, D]$ | batch | ("data", None) | $\left[ \frac{B}{X}, D \right]$ |
| $W_{Q}, W_{K}, W_{V}$ | $[D, N*H]$ | dense_in | ("data", None) | $\left[ \frac{D}{X}, N*H \right]$ |
| $W_{O}$ | $[N*H, D]$ | dense_out | (None, "data") | $\left[ N*H, \frac{D}{X} \right]$ |
| $W_{1}, W_{3}$ | $[D, F]$ | dense_in | ("data", None) | $\left[ \frac{D}{X}, F \right]$ |
| $W_{2}$ | $[F, D]$ | dense_out | (None, "data") | $\left[ F, \frac{D}{X} \right]$ |
| $\text{RMSNorm}$ | $[D]$ | norm | (None) | $[D]$ |
| $\text{Embedding}$ | $[V, D]$ | embedding | (None, None) | $[V, D]$ |
| $\text{LM Head}$ | $[D, V]$ | lm_head | (None, None) | $[D, V]$ |
And here’s what it would look like in code:
def get_sharding_config_for_mode(mode: str) -> ShardingConfig:
...
if mode == "fsdp":
return ShardingConfig(
dense_in=("data", None),
dense_out=(None, "data"),
embedding=(None, None),
lm_head=(None, None),
norm=(None,),
)
...TP
In TP, we have the following:
\[\text{In}[B, D] \cdot_D W_{1}[D, F_Y] \cdot_F W_{2}[F_Y, D] \rightarrow \text{Out}[B, D]\]where we want to shard our weights along the “tensor” axis.
| Component | Shape | Sharding Name | TP | Resulting Shape (per device) |
|---|---|---|---|---|
| $\text{Input}$ | $[B, D]$ | batch | (None, None) | $[B, D]$ |
| $W_{Q}, W_{K}, W_{V}$ | $[D, N*H]$ | dense_in | (None, "tensor") | $\left[ D, \frac{N}{Y}*H \right]$ |
| $W_{O}$ | $[N*H, D]$ | dense_out | ("tensor", None) | $\left[ \frac{N}{Y}*H, D \right]$ |
| $W_{1}, W_{3}$ | $[D, F]$ | dense_in | (None, "tensor") | $\left[ D, \frac{F}{Y} \right]$ |
| $W_{2}$ | $[F, D]$ | dense_out | ("tensor", None) | $\left[ \frac{F}{Y}, D \right]$ |
| $\text{RMSNorm}$ | $[D]$ | norm | (None) | $[D]$ |
| $\text{Embedding}$ | $[V, D]$ | embedding | (None, "tensor") | $\left[ V, \frac{D}{Y} \right]$ |
| $\text{LM Head}$ | $[D, V]$ | lm_head | ("tensor", None) | $\left[ \frac{D}{Y}, V \right]$ |
And here’s what the code would look like:
def get_sharding_config_for_mode(mode: str) -> ShardingConfig:
...
if mode == "tp":
return ShardingConfig(
dense_in=(None, "tensor"),
dense_out=("tensor", None),
embedding=(None, "tensor"),
lm_head=("tensor", None),
norm=(None,),
)
...FSDP+TP
Finally, in FSDP+TP, we have the following:
\[\text{In}[B_X, D_Y] \cdot_D W_{1}[D_X, F_Y] \cdot_F W_{2}[F_Y, D_X] \rightarrow \text{Out}[B_X, D_Y]\]| Component | Shape | Sharding Name | FSDP+TP | Resulting Shape (per device) |
|---|---|---|---|---|
| $\text{Input}$ | $[B, D]$ | batch | ("data", None) | $\left[ \frac{B}{X}, D \right]$ |
| $W_{Q}, W_{K}, W_{V}$ | $[D, N*H]$ | dense_in | ("data", "tensor") | $\left[ \frac{D}{X}, \frac{N}{Y}*H \right]$ |
| $W_{O}$ | $[N*H, D]$ | dense_out | ("tensor", "data") | $\left[ \frac{N}{Y}*H, \frac{D}{X} \right]$ |
| $W_{1}, W_{3}$ | $[D, F]$ | dense_in | ("data", "tensor") | $\left[ \frac{D}{X}, \frac{F}{Y} \right]$ |
| $W_{2}$ | $[F, D]$ | dense_out | ("tensor", "data") | $\left[ \frac{F}{Y}, \frac{D}{X} \right]$ |
| $\text{RMSNorm}$ | $[D]$ | norm | (None) | $[D]$ |
| $\text{Embedding}$ | $[V, D]$ | embedding | (None, "tensor") | $\left[ V, \frac{D}{Y} \right]$ |
| $\text{LM Head}$ | $[D, V]$ | lm_head | ("tensor", None) | $\left[ \frac{D}{Y}, V \right]$ |
And here’s what the code looks like:
def get_sharding_config_for_mode(mode: str) -> ShardingConfig:
...
if mode == "fsdp_tp":
return ShardingConfig(
dense_in=("data", "tensor"),
dense_out=("tensor", "data"),
embedding=(None, "tensor"),
lm_head=("tensor", None),
norm=(None,),
)
...As you can see, implementing distributed training at this point is mostly plumbing: 1) initialize the model inside a mesh scope, 2) place the external token batch with the appropriate batch sharding, and 3) let JAX/XLA infer the collective communication implied by those layouts. That’s the main appeal of JAX’s sharding model—rather than manually writing all-gathers and reduce-scatters at the Python level, we can simply describe the layout we want and let the compiler handle these communications implicitly.
We also provide a PyTorch implementation of these distributed training methods (DP, TP, FSDP, FSDP+TP) for comparison. You can find it here. Note how much more complex it is!
Conclusion
Hopefully this was helpful! In this blog post, we went over some of the basic JAX/NNX concepts, demonstrated how we can build a language model from scratch using NNX, and discussed how we can implement distributed training.
Now, you can try applying your knowledge with our assignment or refer to our reference implementations.
This blog is still a work in progress! Some of the things that we plan to release soon:
- Introduction to Roofline Analysis
- Empirical Results and Comparison
- Mixture of Experts (MoE) implementation
- Additional LLM Tricks
We aim for this to be an easily customizable and useful resource for tinkering around in JAX.
Acknowledgements
Thank you to Henry Ko for providing helpful feedback on this blog post! We would also like to thank Machine Learning @ Berkeley for providing compute resources and feedback on an early version.
References
Austin et al., “How to Scale Your Model”, Google DeepMind, online, 2025.
Hashimoto et al., “CS336: Language Modeling from Scratch”, Stanford NLP, online, 2026.
https://pytorch.org/blog/overview-of-pytorch-autograd-engine/
https://docs.jax.dev/en/latest/
https://flax.readthedocs.io/en/stable/
Enjoy Reading This Article?
Here are some more articles you might like to read next: